









— or sign up using email —
 See how it works
See how it works
| Categories | Rank | Trend in last 3 yrs |
| Structural Biology | #21 of 48 |




Impact Factor |
CiteRatio |
||||||||||||||||||||||
Determines the importance of a journal by taking a measure of frequency with which the average article in a journal has been cited in a particular year. |
A measure of average citations received per peer-reviewed paper published in the journal. |
||||||||||||||||||||||
3.07118% from 2018
|
5.518% from 2019
|
||||||||||||||||||||||
|
|
SCImago Journal Rank (SJR) |
Source Normalized Impact per Paper (SNIP) |
||||||||||||||||||||||||
Measures weighted citations received by the journal. Citation weighting depends on the categories and prestige of the citing journal. |
Measures actual citations received relative to citations expected for the journal's category. |
||||||||||||||||||||||||
2.56430% from 2019
|
1.0184% from 2019
|
||||||||||||||||||||||||
|
|

Guideline source: View
All company, product and service names used in this website are for identification purposes only. All product names, trademarks and registered trademarks are property of their respective owners.
Use of these names, trademarks and brands does not imply endorsement or affiliation. Disclaimer Notice

Elsevier
The Journal of Structural Biology publishes papers dealing with the structural analysis of living material at every level of organization by all methods that lead to an understanding of biological function in terms of molecular and supermolecular structure. Techniques covered ...... Read More
| [25] | |
| G. E. Blonder, M. Tinkham, T. M. Klapwijk, Transition from metallic to tunneling regimes in superconducting microconstrictions: Excess current, charge imbalance, and supercurrent conversion, Phys. Rev. B 25 (7) (1982) 4515–4532. URL 10.1103/PhysRevB.25.4515 |

SciSpace is a very innovative solution to the formatting problem and existing providers, such as Mendeley or Word did not really evolve in recent years.
(Before submission check for plagiarism via Turnitin)
 Less than 3 minutes
Less than 3 minutes
With SciSpace, you do not need a word template for Journal of Structural Biology.
It automatically formats your research paper to Elsevier formatting guidelines and citation style.
You can download a submission ready research paper in pdf, LaTeX and docx formats.

Time taken to format a paper and Compliance with guidelines
SciSpace has partnered with Turnitin, the leading provider of Plagiarism Check software.
Using this service, researchers can compare submissions against more than 170 million scholarly articles, a database of 70+ billion current and archived web pages. How Turnitin Integration works?


One editor, 100K journal formats – world's largest collection of journal templates
With such a huge verified library, what you need is already there.
Journal of Structural Biology format uses elsarticle-num citation style.
Automatically format and order your citations and bibliography in a click.
SciSpace allows imports from all reference managers like Mendeley, Zotero, Endnote, Google Scholar etc.
Absolutely not! Our tool has been designed to help you focus on writing. You can write your entire paper as per the Journal of Structural Biology guidelines and auto format it.
Yes, the template is compliant with the Journal of Structural Biology guidelines. Our experts at SciSpace ensure that. If there are any changes to the journal's guidelines, we'll change our algorithm accordingly.
Of course! We support all the top citation styles, such as APA style, MLA style, Vancouver style, Harvard style, and Chicago style. For example, when you write your paper and hit autoformat, our system will automatically update your article as per the Journal of Structural Biology citation style.
Sign up for our free trial, and you'll be able to use all our features for seven days. You'll see how helpful they are and how inexpensive they are compared to other options, Especially for Journal of Structural Biology.
Yes. You can choose the right template, copy-paste the contents from the word document, and click on auto-format. Once you're done, you'll have a publish-ready paper Journal of Structural Biology that you can download at the end.
It only takes a matter of seconds to edit your manuscript. Besides that, our intuitive editor saves you from writing and formatting it in Journal of Structural Biology.
It is possible to find the Word template for any journal on Google. However, why use a template when you can write your entire manuscript on SciSpace , auto format it as per Journal of Structural Biology's guidelines and download the same in Word, PDF and LaTeX formats? Give us a try!.
Of course! You can do this using our intuitive editor. It's very easy. If you need help, our support team is always ready to assist you.
SciSpace's Journal of Structural Biology is currently available as an online tool. We're developing a desktop version, too. You can request (or upvote) any features that you think would be helpful for you and other researchers in the "feature request" section of your account once you've signed up with us.
Sure. You can request any template and we'll have it setup within a few days. You can find the request box in Journal Gallery on the right side bar under the heading, "Couldn't find the format you were looking for like Journal of Structural Biology?”
After writing your paper autoformatting in Journal of Structural Biology, you can download it in multiple formats, viz., PDF, Docx, and LaTeX.
To be honest, the answer is no. The impact factor is one of the many elements that determine the quality of a journal. Few of these factors include review board, rejection rates, frequency of inclusion in indexes, and Eigenfactor. You need to assess all these factors before you make your final call.

We extracted this data from Sherpa Romeo to help researchers understand the access level of this journal in accordance with the Sherpa Romeo Archiving Policy for Journal of Structural Biology. The table below indicates the level of access a journal has as per Sherpa Romeo's archiving policy.
| RoMEO Colour | Archiving policy |
| Green | Can archive pre-print and post-print or publisher's version/PDF |
| Blue | Can archive post-print (ie final draft post-refereeing) or publisher's version/PDF |
| Yellow | Can archive pre-print (ie pre-refereeing) |
| White | Archiving not formally supported |
The 5 most common citation types in order of usage for Journal of Structural Biology are:.
| S. No. | Citation Style Type |
| 1. | Author Year |
| 2. | Numbered |
| 3. | Numbered (Superscripted) |
| 4. | Author Year (Cited Pages) |
| 5. | Footnote |
It is possible to find the Word template for any journal on Google. However, why use a template when you can write your entire manuscript on SciSpace , auto format it as per Journal of Structural Biology's guidelines and download the same in Word, PDF and LaTeX formats? Give us a try!.
Yes, SciSpace provides this functionality. After signing up, you would need to import your existing references from Word or Bib file to SciSpace. Then SciSpace would allow you to download your references in Journal of Structural Biology Endnote style according to Elsevier guidelines.
Instant formatting to 100% publisher guidelines on - SciSpace.
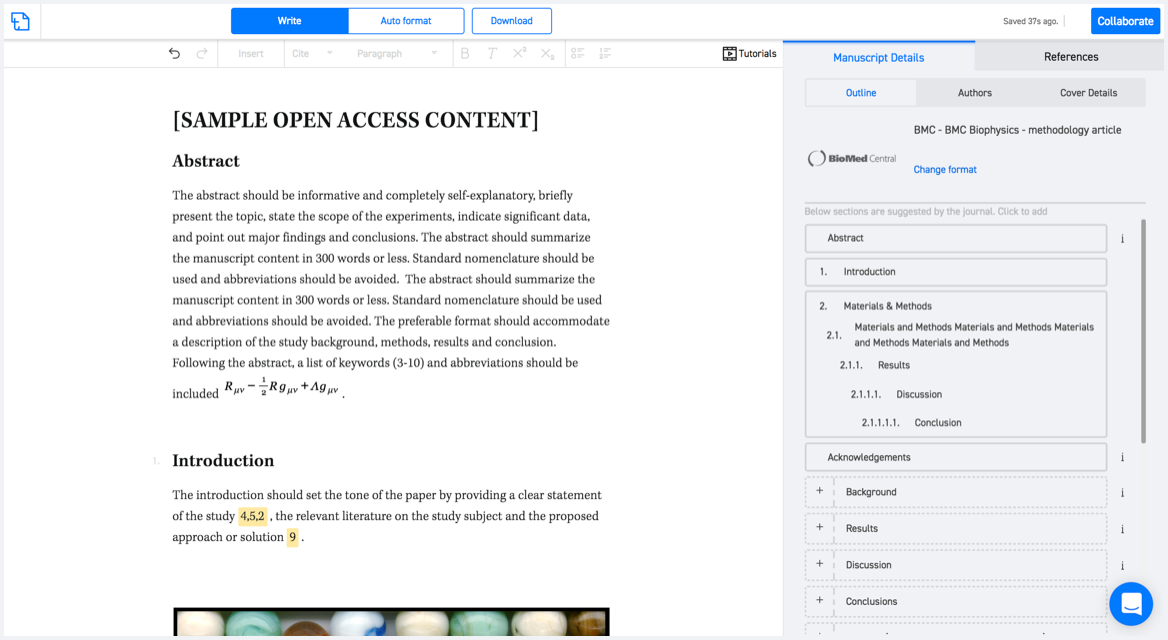

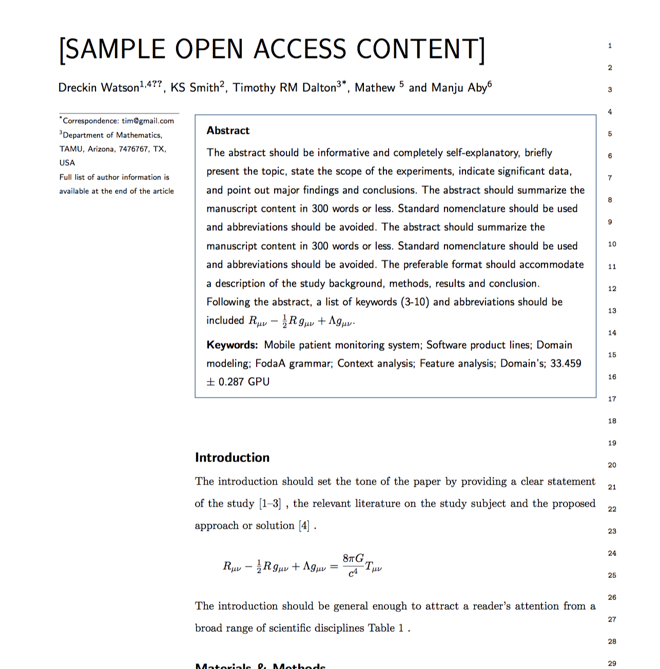
Typset automatically formats your research paper to Journal of Structural Biology formatting guidelines and citation style.


 Verifed journal formats
Verifed journal formatsOne editor, 100K journal formats.
With the largest collection of verified journal formats, what you need is already there.

I spent hours with MS word for reformatting. It was frustrating - plain and simple. With SciSpace, I can draft my manuscripts and once it is finished I can just submit. In case, I have to submit to another journal it is really just a button click instead of an afternoon of reformatting.
