



read more
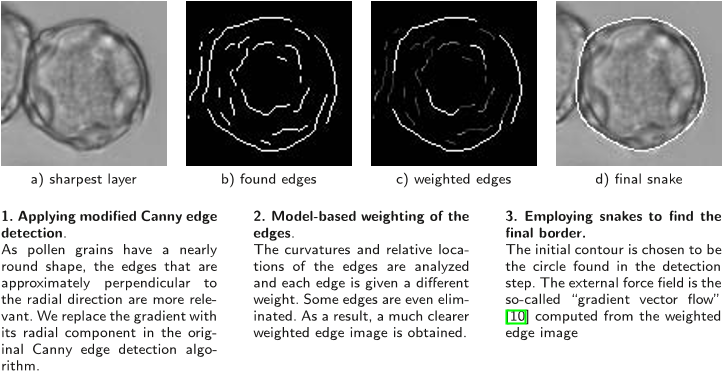








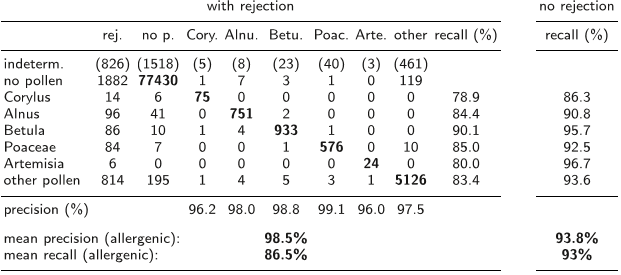

Due to the sparsity of non-zero-values in the synthetic channels the resulting integral features are highly localized in the real space, while the framework automatically guarantees the desired invariance properties.
A few such pollen grains per m3 of air can already cause allergic reactions) the avoidance of false positives is one of the most important requirements for a fully automated system.
While the authors achieve full invariance to radial deformations by full Haar-integration the authors can only reach robustness to local deformations by partial Haar-integration.
The first step in processing the pollen monitor data set is the detection of circular objects with voxel-wise vector based gray-scale invariants, similar to those in [8].
From the training set only the “clean” (not agglomerated, not contaminated) pollen and the “non-pollen” particles from a few samples were used to train the support vector machine (SVM) using the RBF-kernel (radial basis function) and the one-vs-rest multi-class approach.
For the application on the pollen monitor data set (rotational invariance only around the z-axis), q is split into a radial distance qr to the segmentation border and the z-distance to the central plane qz.
The best sampling of the parameter space of the kernel functions (corresponding to the inner class deformations of the objects), was found by cross validation on the training data set, resulting in Nqr ×Nqz ×Nc×n = 31×11×16×16 = 87296 “structural” features (using kernel function k1) and 8 “shape” features (usingkernel function k2).
Within the BMBF-founded project “OMNIBUSS” a first demonstrator of a fully automated online pollen monitor was developed, that integrates the collection, preparation and microscopic analysis of air samples.
This is achieved by creating synthetic channels containing the segmentation borders and employing special parameterized kernel functions.
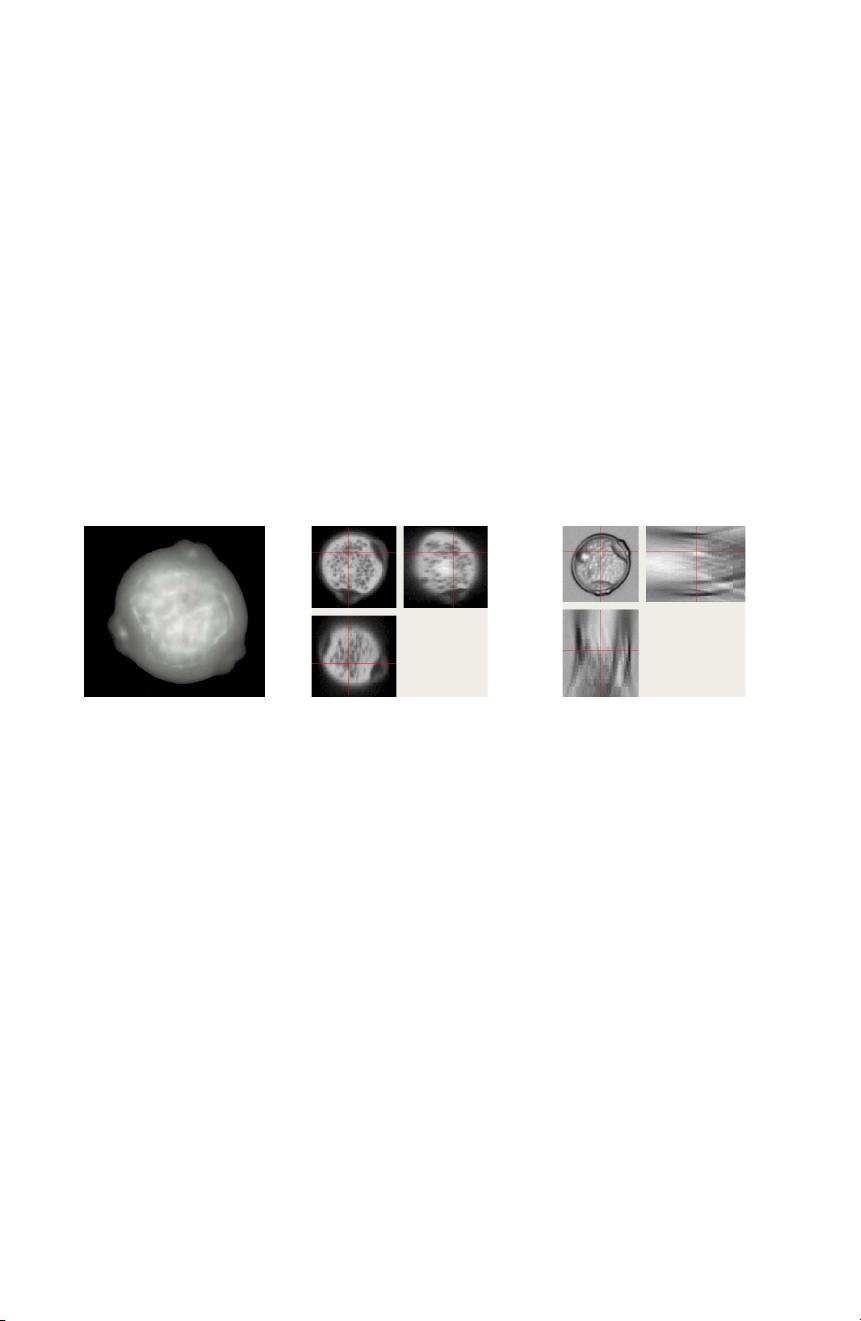
The “pollen monitor data set” contains about 180,000 airborne particles including about 22,700 pollen grains from air samples that were collected, preparedand recorded with transmitted light microscopy from the online pollen monitor from March to September 2006 in Freiburg and Zürich (fig. 1c).
the group of arbitrary deformations GD and the group of rotations GR the final Haar integral becomes:T = ∫GR∫Gγ∫GDf ( gRgγgDS, gRgγgDX ) p(D) dgD dgγ dgR , (8)where p(D) is the probability for the occurrence of the local displacement field D. The transformation of the data set is described by (gX)(x) =: X(x′), wherex′ = Rx︸︷︷︸ rotation + γ(Rx)︸ ︷︷ ︸ global deformation+
For 3D rotations this framework uses a spherical-harmonics series expansion, and for planar rotations around the z-axis it is simplified to a Fourier series expansion.
As most pollen grains are nearly spherical and the subtle differences are mainly found near the surface, a pollen expert needs the full 3D information (usually by “focussing through” the transparent pollen grain).