
All figures (9)
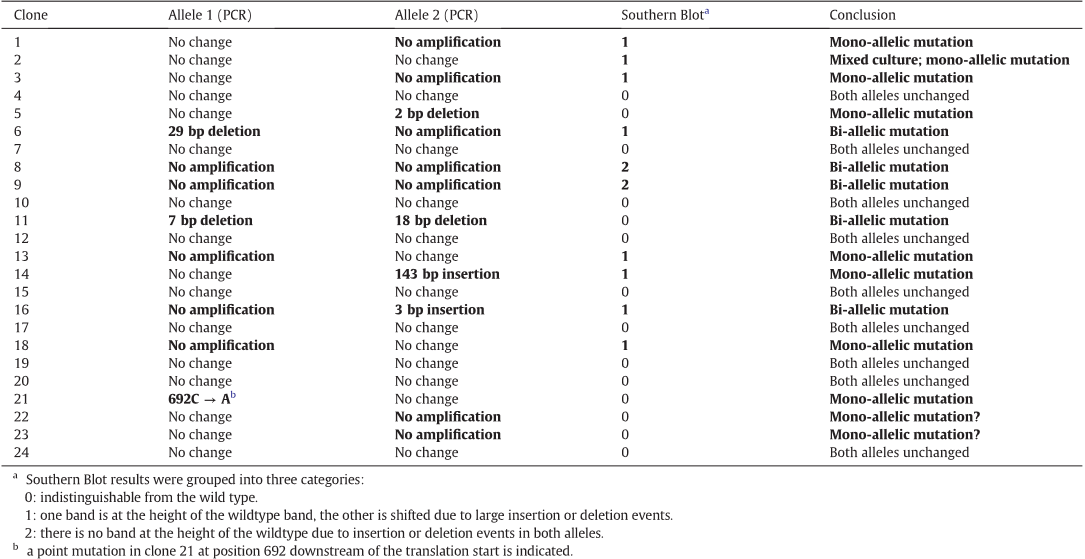 Table 1 Evaluation of the screening of the two alleles via PCR and Southern Blot. Detected mutagenic events are indicated in bold.
Table 1 Evaluation of the screening of the two alleles via PCR and Southern Blot. Detected mutagenic events are indicated in bold. Fig. 4. Southern Blots using digested genomic DNA of either wild type or transformed cell lines. Shifts of the DNA fragments can be caused either by insertion or deletion events. Shifted DNA bands due to insertions or deletions are marked with arrows.
Fig. 4. Southern Blots using digested genomic DNA of either wild type or transformed cell lines. Shifts of the DNA fragments can be caused either by insertion or deletion events. Shifted DNA bands due to insertions or deletions are marked with arrows. Fig. 5. Southern Blot using digested genomicDNA of eitherwild type or transformed cell lines. Shifts of theDNA fragments can be caused either by insertion or deletion events. Shifted DNA bands due to insertions and deletions are marked with arrows.
Fig. 5. Southern Blot using digested genomicDNA of eitherwild type or transformed cell lines. Shifts of theDNA fragments can be caused either by insertion or deletion events. Shifted DNA bands due to insertions and deletions are marked with arrows.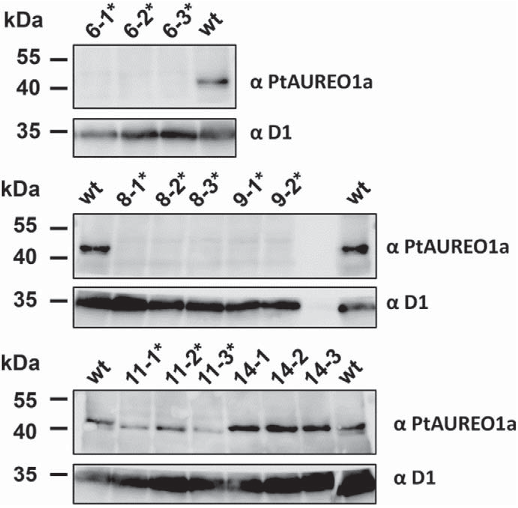 Fig. 6. Western Blot of wild type and two mono-allelic as well as four bi-allelic cell lines (marked with a *) after re-isolation using the PtAUREO1a antiserum. The expected molecular weight of PtAUREO is 41.5 kDa. A D1-specific antiserum was used as a loading control.
Fig. 6. Western Blot of wild type and two mono-allelic as well as four bi-allelic cell lines (marked with a *) after re-isolation using the PtAUREO1a antiserum. The expected molecular weight of PtAUREO is 41.5 kDa. A D1-specific antiserum was used as a loading control.![Fig. 1. Schematic overview of the TALEN assembly process (according to [33]) and the resulting plasmids. Six monomers are assembled into hexamers corresponding to the desired target site in a golden-gate type reaction from a library consisting of 72 monomers (18 positions of the TALEN target site and 4 different RVDs), allowing assembly of multiple fragments in the desired order in a single step. Three of these hexamers are then ligated in a second golden gate-type reaction into the plasmid containing the first RVD (always NG) and the last half RVD (HD, NG, NI or NN, depending on the target site). Correct insertion of the target sequence needs to be verified by colony PCR (insert size: 2.2 kbp). Clones indicated to be positive by colony PCR should be verified by restriction digest with AfeI (expected fragment lengths: 5 kbp, 2.2 kbp and 165 bp; exemplary shown in Fig. S1) as well as sequencing of the inserted fragment. fcpA: FcpA ( = Lhcf1) promoter; FokI: endonuclease; N-/C-term: N and C terminus, respectively; Nat: nourseothricin resistance cassette; NR: nitrate reductase promoter; RVD: repeat variable di-residue; Sh ble: Zeocin resistance cassette.](/figures/fig-1-schematic-overview-of-the-talen-assembly-process-lkguikzl.png) Fig. 1. Schematic overview of the TALEN assembly process (according to [33]) and the resulting plasmids. Six monomers are assembled into hexamers corresponding to the desired target site in a golden-gate type reaction from a library consisting of 72 monomers (18 positions of the TALEN target site and 4 different RVDs), allowing assembly of multiple fragments in the desired order in a single step. Three of these hexamers are then ligated in a second golden gate-type reaction into the plasmid containing the first RVD (always NG) and the last half RVD (HD, NG, NI or NN, depending on the target site). Correct insertion of the target sequence needs to be verified by colony PCR (insert size: 2.2 kbp). Clones indicated to be positive by colony PCR should be verified by restriction digest with AfeI (expected fragment lengths: 5 kbp, 2.2 kbp and 165 bp; exemplary shown in Fig. S1) as well as sequencing of the inserted fragment. fcpA: FcpA ( = Lhcf1) promoter; FokI: endonuclease; N-/C-term: N and C terminus, respectively; Nat: nourseothricin resistance cassette; NR: nitrate reductase promoter; RVD: repeat variable di-residue; Sh ble: Zeocin resistance cassette.
Fig. 1. Schematic overview of the TALEN assembly process (according to [33]) and the resulting plasmids. Six monomers are assembled into hexamers corresponding to the desired target site in a golden-gate type reaction from a library consisting of 72 monomers (18 positions of the TALEN target site and 4 different RVDs), allowing assembly of multiple fragments in the desired order in a single step. Three of these hexamers are then ligated in a second golden gate-type reaction into the plasmid containing the first RVD (always NG) and the last half RVD (HD, NG, NI or NN, depending on the target site). Correct insertion of the target sequence needs to be verified by colony PCR (insert size: 2.2 kbp). Clones indicated to be positive by colony PCR should be verified by restriction digest with AfeI (expected fragment lengths: 5 kbp, 2.2 kbp and 165 bp; exemplary shown in Fig. S1) as well as sequencing of the inserted fragment. fcpA: FcpA ( = Lhcf1) promoter; FokI: endonuclease; N-/C-term: N and C terminus, respectively; Nat: nourseothricin resistance cassette; NR: nitrate reductase promoter; RVD: repeat variable di-residue; Sh ble: Zeocin resistance cassette. Table 2 Analysis of pigment composition and non-photochemical quenching (NPQ) capacity of wild type (wt) and the four bi-allelic PtAUREO1a knockout strains (6, 8, 9 and 11). Chlorophyll a (Chl a) was determined photometrically (wild type n=5, bi-allelicmutants n=3), whereas the other pigmentswere assayed by HPLC (n=2). Pigmentswere isolated from strains being inmid exponential growth phase and having comparable Chl a per culture-volume ratios. NPQwasmeasured by PAM fluorometry (n=3). Statistical significant differences compared to the wild type were calculated using Student's t-test (*: p b 0.01; **:p b 0.001). XC pool: xanthopyll cycle pool (diadinoxanthin + diatoxanthin).
Table 2 Analysis of pigment composition and non-photochemical quenching (NPQ) capacity of wild type (wt) and the four bi-allelic PtAUREO1a knockout strains (6, 8, 9 and 11). Chlorophyll a (Chl a) was determined photometrically (wild type n=5, bi-allelicmutants n=3), whereas the other pigmentswere assayed by HPLC (n=2). Pigmentswere isolated from strains being inmid exponential growth phase and having comparable Chl a per culture-volume ratios. NPQwasmeasured by PAM fluorometry (n=3). Statistical significant differences compared to the wild type were calculated using Student's t-test (*: p b 0.01; **:p b 0.001). XC pool: xanthopyll cycle pool (diadinoxanthin + diatoxanthin).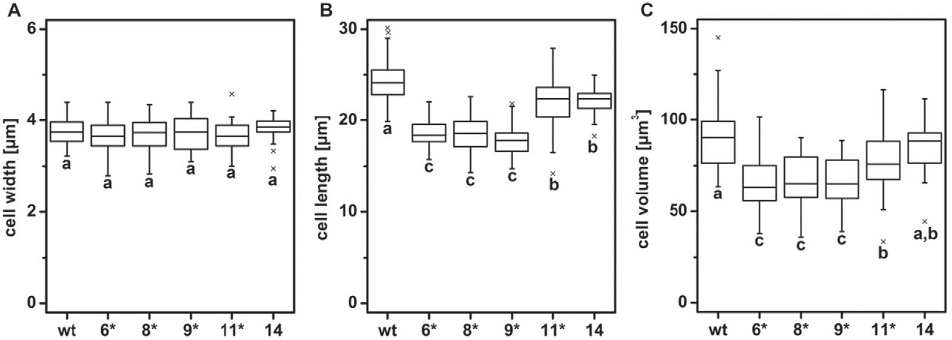 Fig. 7. 50 cells of the wildtype and eachmutant strain (6, 8, 9, 11, 14) were examinedmicroscopically and cell width (A) and length (B) wasmeasured. Cell volume was approximated as two cone shapes (C). The bi-allelic knockout mutants are marked with a *. Whiskers represent the outermost data point within the 1.5-fold of the interquartile range; outliers are represented by an x. Different letters represent statistical significant differences in cell width, length or volume between strains (One-way ANOVA, followed by Tukey's HSD, p b 0.01).
Fig. 7. 50 cells of the wildtype and eachmutant strain (6, 8, 9, 11, 14) were examinedmicroscopically and cell width (A) and length (B) wasmeasured. Cell volume was approximated as two cone shapes (C). The bi-allelic knockout mutants are marked with a *. Whiskers represent the outermost data point within the 1.5-fold of the interquartile range; outliers are represented by an x. Different letters represent statistical significant differences in cell width, length or volume between strains (One-way ANOVA, followed by Tukey's HSD, p b 0.01). Fig. 3. Schematic drawing of PtAureo1a and its geneproduct PtAUREO1a including conserved domains (bZIP in red and LOV inblue). The TALEN recognition sites are indicated by a grey line and the FokI endonuclease domains are symbolized by scissors. The predicted cutting site of the TALEN pair within the gene and its relative location within the gene product and its conserved domains is indicated by dashed lines. The allele-specific difference of PtAureo1a at position 81 (T/G) in strain Pt4 is indicated by an arrow, an 102 bp intron region by an inverted triangle shape and the binding site of the Southern blot probe by a magenta-colored line.
Fig. 3. Schematic drawing of PtAureo1a and its geneproduct PtAUREO1a including conserved domains (bZIP in red and LOV inblue). The TALEN recognition sites are indicated by a grey line and the FokI endonuclease domains are symbolized by scissors. The predicted cutting site of the TALEN pair within the gene and its relative location within the gene product and its conserved domains is indicated by dashed lines. The allele-specific difference of PtAureo1a at position 81 (T/G) in strain Pt4 is indicated by an arrow, an 102 bp intron region by an inverted triangle shape and the binding site of the Southern blot probe by a magenta-colored line. Fig. 2.Workflow and recommendations for design and assembly of TALEN constructs, as well as screening of the obtained transformants. An estimated timeframe is given for each step.
Fig. 2.Workflow and recommendations for design and assembly of TALEN constructs, as well as screening of the obtained transformants. An estimated timeframe is given for each step.





![Fig. 1. Schematic overview of the TALEN assembly process (according to [33]) and the resulting plasmids. Six monomers are assembled into hexamers corresponding to the desired target site in a golden-gate type reaction from a library consisting of 72 monomers (18 positions of the TALEN target site and 4 different RVDs), allowing assembly of multiple fragments in the desired order in a single step. Three of these hexamers are then ligated in a second golden gate-type reaction into the plasmid containing the first RVD (always NG) and the last half RVD (HD, NG, NI or NN, depending on the target site). Correct insertion of the target sequence needs to be verified by colony PCR (insert size: 2.2 kbp). Clones indicated to be positive by colony PCR should be verified by restriction digest with AfeI (expected fragment lengths: 5 kbp, 2.2 kbp and 165 bp; exemplary shown in Fig. S1) as well as sequencing of the inserted fragment. fcpA: FcpA ( = Lhcf1) promoter; FokI: endonuclease; N-/C-term: N and C terminus, respectively; Nat: nourseothricin resistance cassette; NR: nitrate reductase promoter; RVD: repeat variable di-residue; Sh ble: Zeocin resistance cassette.](/figures/fig-1-schematic-overview-of-the-talen-assembly-process-lkguikzl.png)



