









24 citations
23 citations
22 citations
21 citations
...…phylogeny, the G+C content of the genome of strain 114T is, at 34.9 mol%, the lowest among all hitherto analysed members of this genus where DNA G+C contents range from 36.6 to 54.7 mol% (Alvarez-Pérez et al., 2013; Choi et al., 2013; Kim et al., 2014; Li et al., 2014; Smet et al., 2014)....
[...]
20 citations
...…a distance matrix to assign the sequences to operational taxonomic units (OTUs) using Mothur v 1.32.1 at 1 and 3 % dissimilarity cutoffs for bacteria and yeasts, respectively, allowing us to perform further analysis with species-level OTUs (Álvarez-Pérez et al. 2013; Jacquemyn et al. 2013a, b)....
[...]
...With regard to the bacteria, 13 bacterial OTUs were found when sequences were clustered at a 1 % dissimilarity cutoff, corresponding to 12 validly named species (Table 3)....
[...]
...…found to be complementary to those of nectarivorous yeasts such as Metschnikowia that quickly deplete glucose in nectar and enrich this floral reward in fructose, which in turn is preferentially used by the Acinetobacter species (Canto et al. 2008; Herrera et al. 2008; Álvarez-Pérez et al. 2013)....
[...]
...Next, the alignments were used to create a distance matrix to assign the sequences to operational taxonomic units (OTUs) using Mothur v 1.32.1 at 1 and 3 % dissimilarity cutoffs for bacteria and yeasts, respectively, allowing us to perform further analysis with species-level OTUs (Álvarez-Pérez et al. 2013; Jacquemyn et al. 2013a, b)....
[...]
...In this respect, the carbon source consumption profiles observed for nectar-inhabiting Acinetobacter species have been found to be complementary to those of nectarivorous yeasts such as Metschnikowia that quickly deplete glucose in nectar and enrich this floral reward in fructose, which in turn is preferentially used by the Acinetobacter species (Canto et al. 2008; Herrera et al. 2008; Álvarez-Pérez et al. 2013)....
[...]
39,110 citations
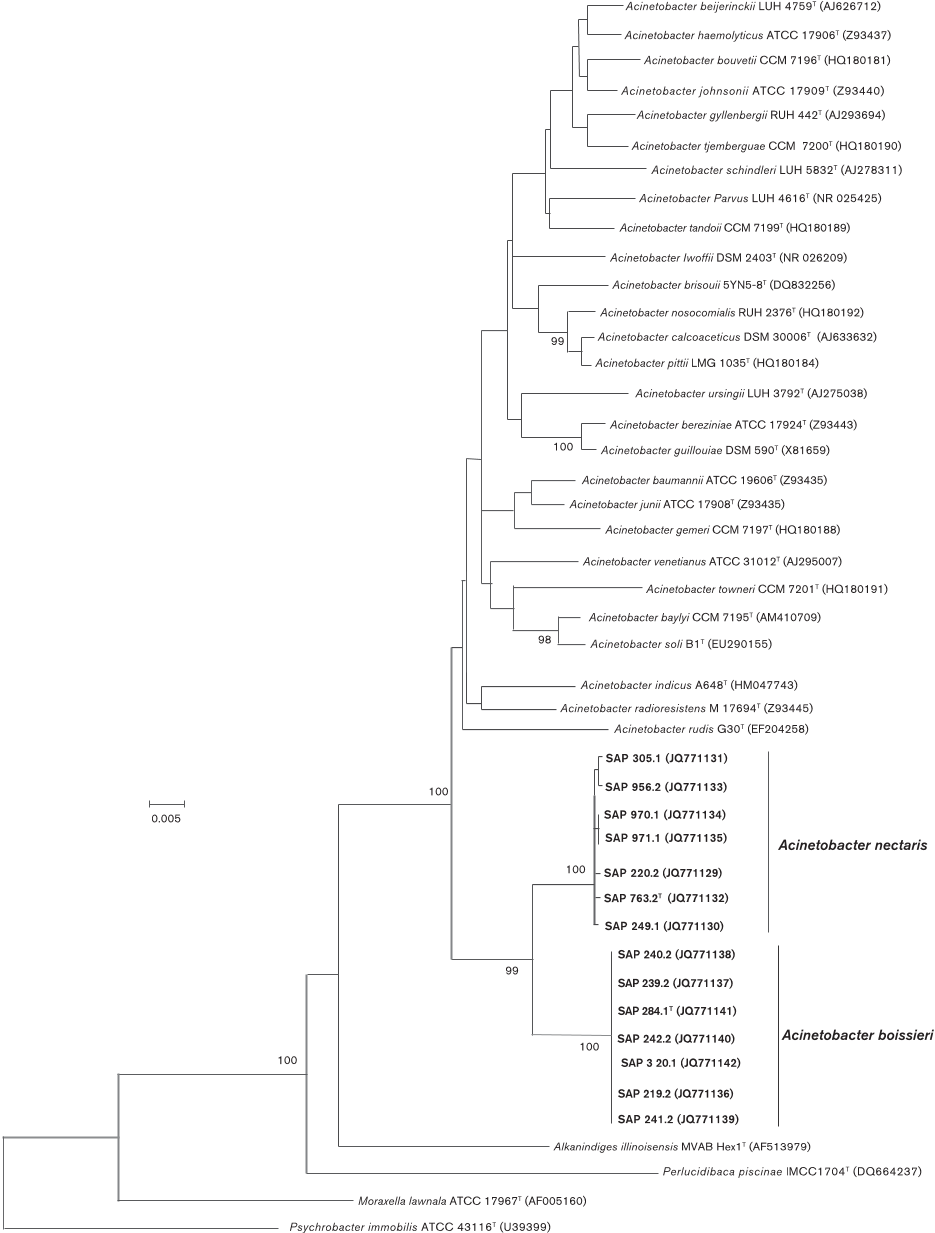
...Following these procedures, 1320 nt positions (98 % of the original alignment) remained for subsequent phylogenetic analysis using the neighbour-joining (NJ) method as implemented in the MEGA 5 software package (Tamura et al., 2011)....
[...]
35,309 citations
8,757 citations
...0 (Hall, 1999) to ensure that all sequences had the same start and end point, and analysed with Gblocks (Castresana, 2000) to eliminate ambiguously aligned regions, using ‘allow gap positions5with half’, ‘minimum length of a block559 and default settings for all other options....
[...]
...…alignment was trimmed with BioEdit v. 7.0.9.0 (Hall, 1999) to ensure that all sequences had the same start and end point, and analysed with Gblocks (Castresana, 2000) to eliminate ambiguously aligned regions, using ‘allow gap positions5with half’, ‘minimum length of a block559 and default settings…...
[...]
...As with the 16S rRNA gene sequence, concatenated sequences of rpoB zones 1 and 2 of the nectar strains and type strains of other species of the genus Acinetobacter were included in a multiple alignment and analysed with Gblocks, which resulted in the selection of 843 nucleotide positions (99 % of the original alignment)....
[...]
...The resulting alignment was trimmed with BioEdit v. 7.0.9.0 (Hall, 1999) to ensure that all sequences had the same start and end point, and analysed with Gblocks (Castresana, 2000) to eliminate ambiguously aligned regions, using ‘allow gap positions5with half’, ‘minimum length of a block559 and default settings for all other options....
[...]
5,300 citations
...The 16S rRNA gene sequences of the novel nectar strains and reference strains of members of the genus Acinetobacter and the family Moraxellaceae were included in a multiple alignment generated by CLUSTAL W (Chenna et al., 2003)....
[...]
4,685 citations
...The DNA was enzymically degraded into nucleosides and the nucleotidic base composition was determined by HPLC, according to the method of Mesbah et al. (1989)....
[...]