




Did you find this useful? Give us your feedback










3 citations
3 citations
3 citations
2 citations
2 citations
13,581 citations
...3 (Excoffier and Lischer, 2010) and the aforementioned population databases were employed to perform pairwise comparisons, Analysis Molecular of Variance (AMOVA), and FST genetic distances were plotted by multidimensional scaling (MDS) with optimum stress of 0....
[...]
...The software Arlequin 3.5.1.3 (Excoffier and Lischer, 2010) and the aforementioned population databases were employed to perform pairwise comparisons, Analysis Molecular of Variance (AMOVA), and FST genetic distances were plotted by multidimensional scaling (MDS) with optimum stress of 0.01 using…...
[...]
7,615 citations
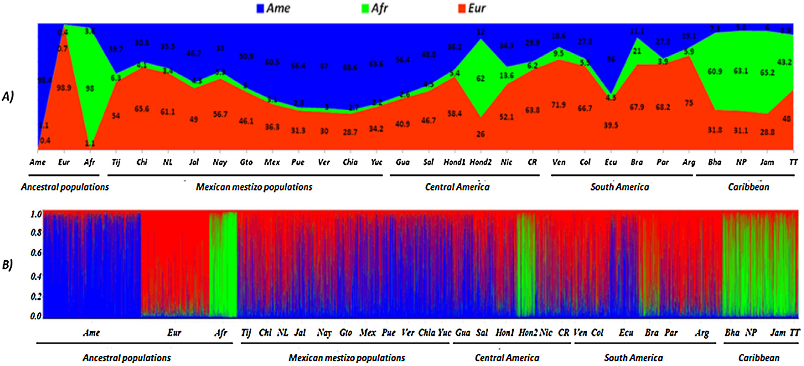
...This is explained by software Leadmix employed before for that purpose (Wang, 2003), whereas in this study the program Structure was used (Falush et al., 2003)....
[...]
...3 (Falush et al., 2003), with a burn-in-period of 10,000 iterations in each parameter and 25 repetitions for each run (K), using the mixture model, allele frequency correlation, and -value separated for populations, with three populations groups identified as the ancestral references (supervised analysis)....
[...]
...components of admixture were estimated in individuals and populations with the software Structure 2.3.3 (Falush et al., 2003), with a burn-in-period of 10,000 iterations in each parameter and 25 repetitions for each run (K), using the mixture model, allele frequency correlation, and -value…...
[...]
1,831 citations
...3 (Excoffier and Lischer, 2010) and the aforementioned population databases were employed to perform pairwise comparisons, Analysis Molecular of Variance (AMOVA), and FST genetic distances were plotted by multidimensional scaling (MDS) with optimum stress of 0.01 using the program SPSS 10.0 for Windows. In addition, genetic distances of Nei (1978) were estimated with the software GDA 1....
[...]
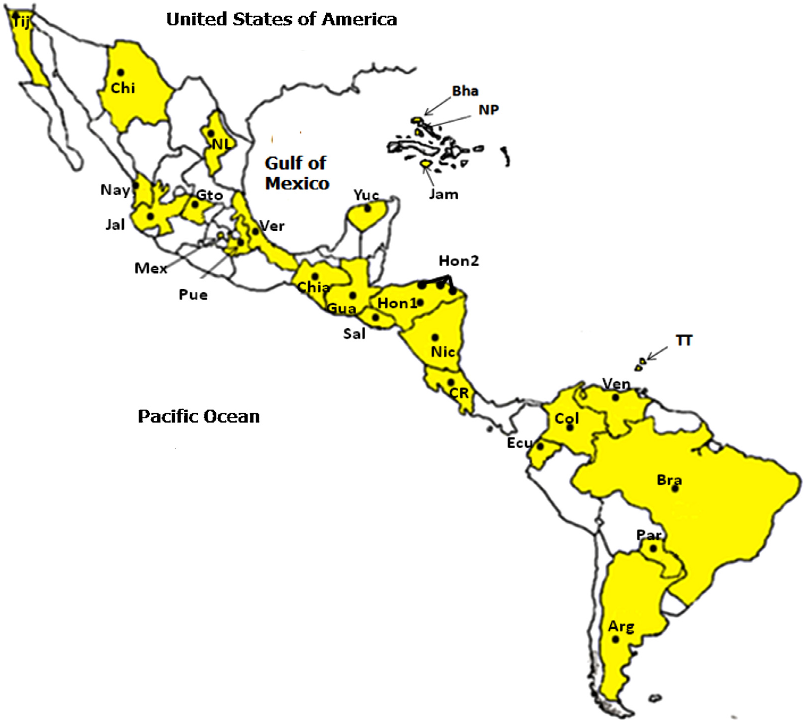
...Different population groups were established considering genetic and geographical criteria using the software SAMOVA 1.0 (Dupanloup et al., 2002)....
[...]
...0 (Dupanloup et al., 2002)....
[...]
769 citations
...However, our preliminary admixture estimates could be helpful in the biomedical area for complex disease analysis (i.e. case–control studies) where population genetic composition and dynamics of the admixture processes should be clearly understood (Parra et al., 1998)....
[...]
625 citations
...The inclusion of CODIS-STRs in commercial human identification kits has increased the number of population databases that can be used in molecular anthropology studies (Butler, 2006)....
[...]
...We amplified 15 STRs markers (D3S1358, TH01, D21S11, D18S51, D5S818, D13S317, D7S820, D16S539, CSF1PO, vWA, D8S1179, TPOX, FGA, D2S1338 and D19S433) as recommended in the PCR AmpFlSTR Identifiler kit (Applied Biosystems, Foster City, CA)....
[...]
...This is due to their elevated heterozygosity, genome abundance, high mutation rate, and simple analysis based on the polymerase chain reaction (PCR) (Butler, 2006)....
[...]