




Did you find this useful? Give us your feedback









4,409 citations
365 citations
327 citations
...Multi-class discriminative sequence feature frameworks have been proposed (Beer & Tavazoie, 2004; Elemento et al, 2007), but these approaches have also been limited to analysis of mutually exclusive classes....
[...]
285 citations
...Studies of individual TFs suggest that binding affinity to cognate motif instances may play a role in distinguishing multi-condition binding sites from tissue-specific sites (Gertz et al, 2013; Mahony et al, 2014)....
[...]
...Our results support the model that high affinity cognate motif instances are a striking feature of multi-conditionally bound sites across a broad range of TFs (Gertz et al, 2013; Mahony et al, 2014)....
[...]
264 citations
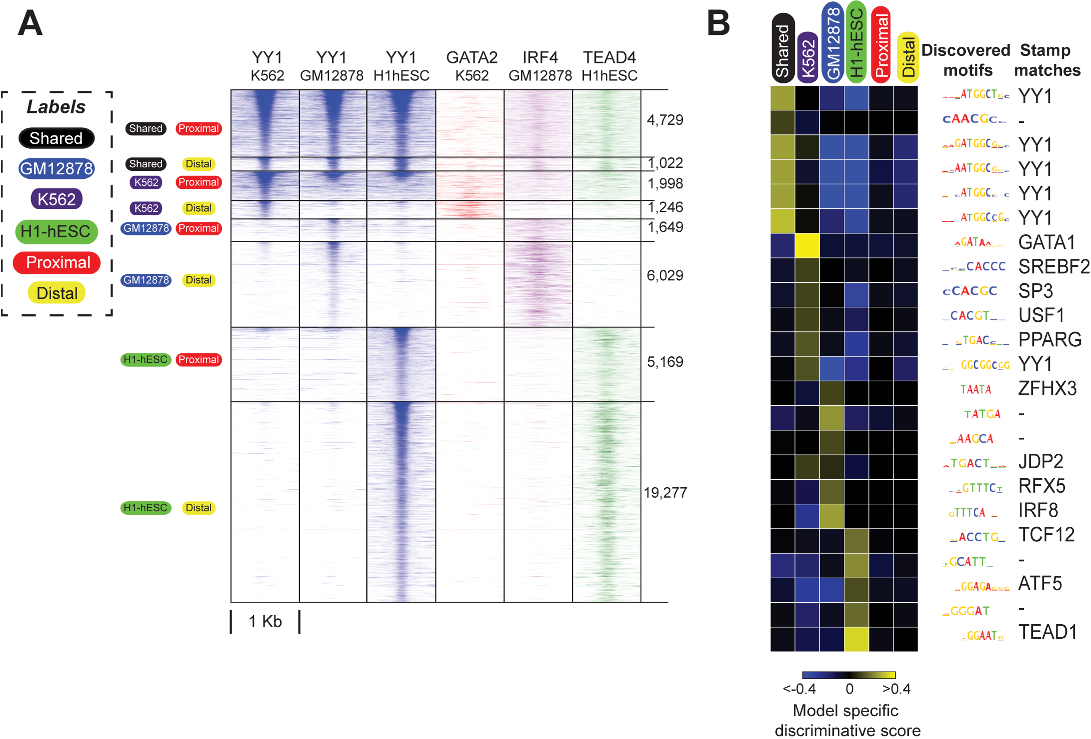
...…a particular TF in multiple cell types (i.e. “shared” or multi-condition sites) are often strongly biased towards being located in gene promoter regions, in contrast to cell-specific binding sites, which are typically distally located (Yip et al, 2012; Wang et al, 2012; Kheradpour & Kellis, 2014)....
[...]
...Similar findings were previously identified at the “high occupancy of transcription-related factors (HOT)” regions (Yip et al, 2012)....
[...]
262 citations
...These results are consistent with the “TF collective” model proposed by Junion and colleagues (Junion et al, 2012)....
[...]
233 citations
...For example (Lee et al, 2011), trained an SVM classifier to discriminate putative enhancers from random sequences using an unbiased set of k-mers as predictors....
[...]
...…convolutional neural networks and support vector machines (SVMs) with various k-mer based sequence kernels (Arvey et al, 2012; Ghandi et al, 2014; Lee et al, 2011), these applications typically focus on discriminating between two mutually exclusive classes (Bailey, 2011; Alipanahi et al, 2015)....
[...]
To calculate collective degree, the authors used a total of 158, 102, and 202 ChIP-seq datasets in GM12878, H1-hESC, and K562 cell-types, respectively.
A significant depletion of motif instances at sites annotated by a label compared to other labels can very likely result in non-positive scores.
Most popular motif-finding methods use unsupervised machinelearning approaches to discover motifs in ‘foreground’ input sequences that are over-represented with respect to a set of ‘background’ sequences (e.g. “bound” vs. “unbound”, respectively) [1,2].
In other words, while the k-mer weight parameters for each subclass are learned directly from the data, the weight parameters for the labels are learned exclusively through the regularization constraint.
By implicitly accounting for the effects of overlapping annotation labels, SeqUnwinder can deconvolve sequence features associated with motor neuron programming dynamics and ES chromatin status.
the authors found IRF and RUNX motifs enriched at GM12878-specific binding sites for 11 and 7 of the 17 examined TFs, respectively.
One advantage of the “hill-finding” approach is that it implicitly takes into account positional relationships between high-scoring k-mers on the genome; short stretches that contain multiple high-scoring k-mers will form larger “hills”.
Several variants of the basic string kernel (e.g. mismatch kernel [35], di-mismatch kernel [4], wild-card kernel [5,35], and gkm-kernel [36]) have been proposed and have been shown to substantially improve the classifier performance.
Since DREME takes only two classes as input: a foreground set and a background set, the authors ran four different DREME runs for each of the four labels.
binding sites showing significantly differential binding in any of the possible 3 pair-wise comparisons were removed from the shared set.
SeqUnwinder’s characterization of cell-specific motif features in collections of DNase-seq datasets may therefore serve as a source of predictive features for efforts that aim to predict cell-specific TF binding from accessibility experimental data alone [39–41].
To speed-up implementation, the authors restrict the unbiased k-mer features to only those k-mers that are present in at least 5% of the hills.
All sites with significantly greater Isl1/Lhx3 ChIP enrichment at 12h compared to 48h (q-value cutoff of<0.01) were labeled as “early”.
the motifs that the authors previously assigned to early or late TF binding behaviors could have been merely associated with ES-active and ES-inactive sites, respectively.
the cognate motif was not specifically predictive of cell-type-specific labels for the examined TFs, with the exception of H1-hESC-specific sites for CEBPB, NRSF and SRF.