



Did you find this useful? Give us your feedback





16 citations
8 citations
1 citations
24,024 citations
3,229 citations
2,973 citations
1,574 citations
1,517 citations
Future experimental work will be conducted to measure the electron transport rate, intracellular ATP concentration, and RuBisCO gene expression across different quinone redox states to strengthen the proposed hypothesis and further refine the model.
Predictions also indicated that the extent of CO2 fixation is dependent on the amount of ATP present, with the quinone redox state acting as a feed-forward signal to the CBB system.
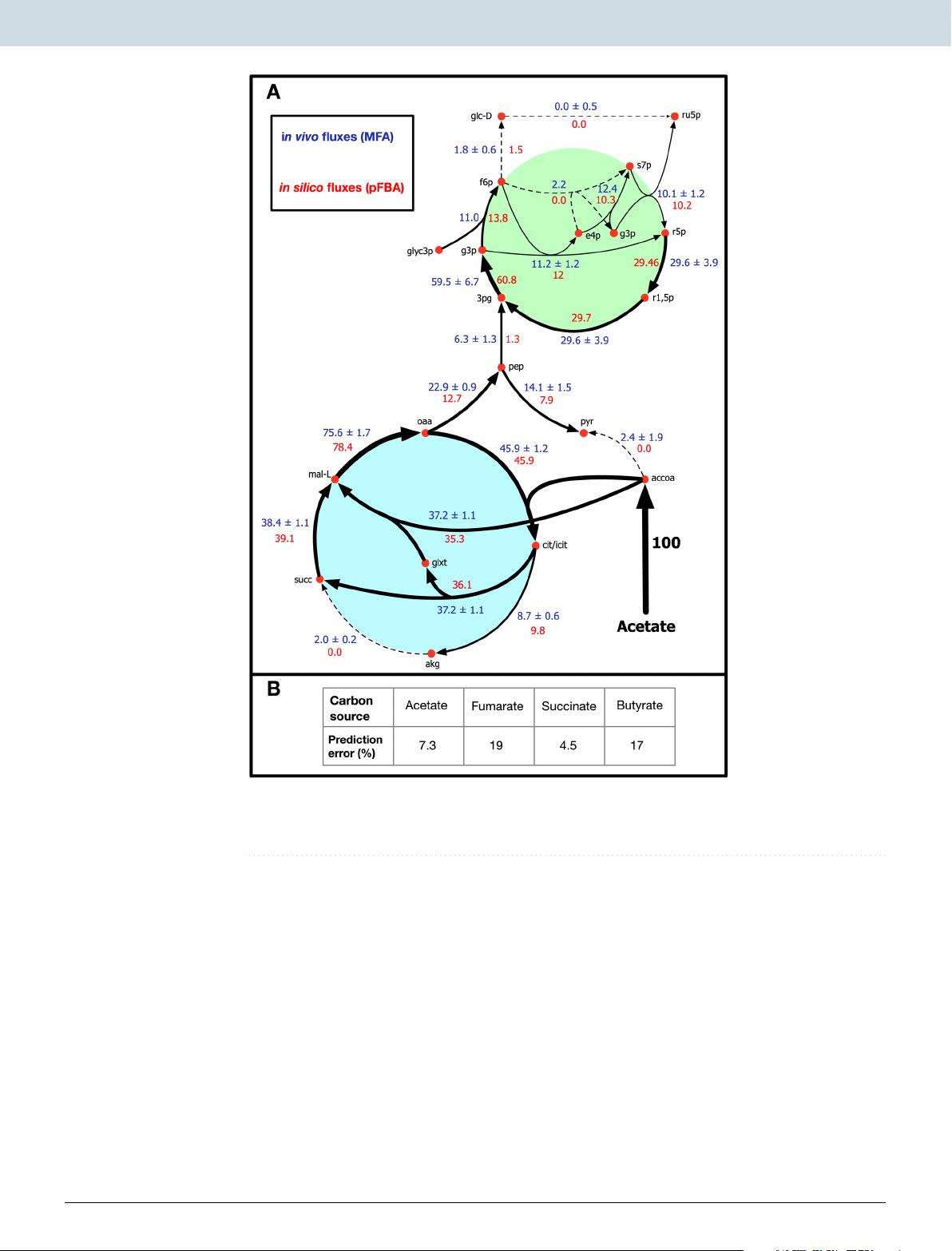
During initial phototrophic growth simulations, growth on any of the four carbon sources (acetate, fumarate, succinate, and butyrate) was observed to be hindered due to the accumulation of excess quinols formed in the TCA cycle.
CO2-fixation using the enzyme ribulose-1,5-biphosphate carboxylase/oxygenase (RuBisCO), nitrogen-fixation through the enzyme nitrogenase12, and supplementation with an electron acceptor (e.g., trimethylamine-N-oxide (TMAO))15 all prevent the inhibitory accumulation of excess reducing agents.
Although the model predicted that the rate of CO2 fixation increased linearly with light uptake rate, kinetic and thermodynamic constrains on the highly inefficient CO2-fixing RuBisCO enzyme50 hinders this process at high light uptake.
Funding to support this work was provided by University of Nebraska-Lincoln Faculty Startup Grant and Nebraska Center for Energy Sciences Research (NCESR) to Rajib Saha.
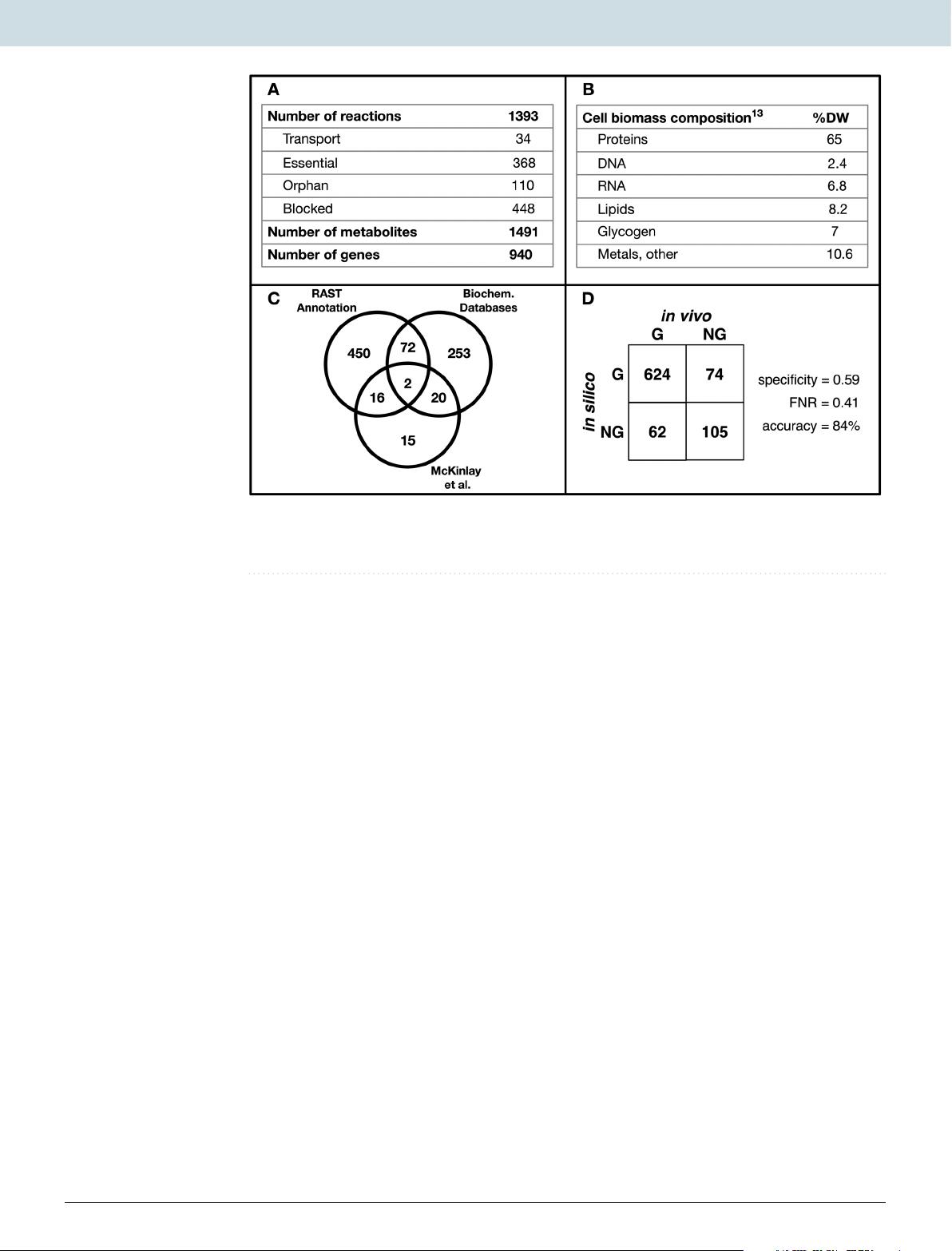
In this study, a genome-scale metabolic network (iRpa940) was used to propose a system-wide mechanistic model of the interactive system that includes photosynthesis, carbon dioxide fixation, and the quinone redox state.
the ModelSEED database33 was used to fill the gaps in the network, and a biomass producing model was generated in KBase32.
Out of the 478 reactions added during gap-filling, 368 were annotated using information from organism-specific databases (see Methods).
Several studies conducted on R. palustris showed that in addition to the Calvin-Benson-Bassham (CBB) cycle’s role of carbon assimilation during autotrophic growth, the pathway plays a major role in maintaining redox balance under heterotrophic conditions10,12–14.
Based on this analysis, it is hypothesized that the quinone redox state acts as a feed-forward controller of the CBB pathway, signaling the amount of ATP available.
the quinone redox state is predicted to act as a feed-forward controller to the energetically expensive CBB pathway, indicating how much ATP is available at a given condition.
After the model indicated the presence of an unidentified quinol sink, in silico simulations were combined with published in vivo flux measurements13,14 to study the effect (and the extent) of the quinone redox state on cellular growth, electron transport rate, and CO2 fixation.
The quinol sink reaction was treated as a parameter in the model and pFBA simulations were conducted at varying quinol oxidation (sink) rates to determine how light uptake (i.e. Electron Transport Rate or ETR), growth, and CO2 fixation are affected by changes in the quinone redox state (Fig. 2).
Flux analysis of the electron transport chain (ETC) revealed that the rate of quinol oxidization through the cytochrome bc1 complex was equivalent to the rate of quinone reduction in the Reaction center (RC).
These results suggest that redox state acts as a feed-forward controller of the highly energy-demanding CBB cycle by regulating the rate of light-generated ATP.