




Did you find this useful? Give us your feedback










79 citations
39 citations
32 citations
...Primary human hepatocytes are still considered to represent a gold standard for hepatic biotransformation studies (Godoy et al. 2018; Gu et al. 2018), whereas HepG2 cells have been reported to represent a useful tool to study the regulation of drug-metabolizing enzymes (Wilkening et al....
[...]
25 citations
...Test strategy/AOP Gu et al., 2018 Relevance of the incubation period in cytotoxicity testing with primary human hepatocytes The publication addresses a central scientific topic of EU-ToxRisk: how long do in vitro experiments need to run to predict human-repeated-dose toxicity?...
[...]
17 citations
...Gu et al. (2018) address a central scientific topic of EU-ToxRisk: How long do in vitro experiments need to run to predict human repeated-dose toxicity? They explored how the variation of the incubation period of a test compound can significantly influence the results of in vitro tests....
[...]
1,988 citations
1,085 citations
706 citations
471 citations
...Currently, primary human hepatocytes represent the gold standard model for in vitro testing of drug metabolism and cytotoxicity (LeCluyse 2001)....
[...]
336 citations
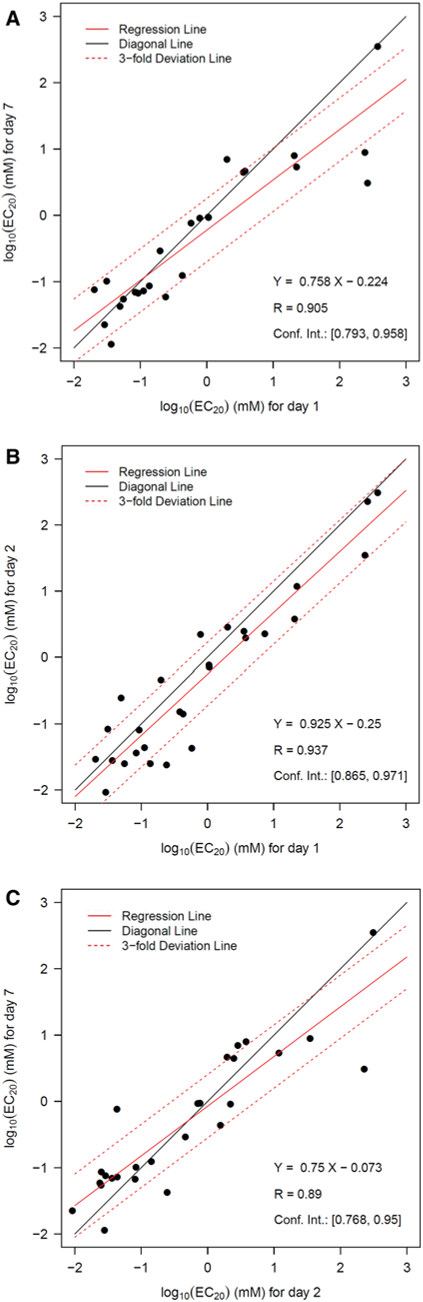
Therefore, 2 days may represent an adequate choice for cytotoxicity tests with human hepatocytes in future studies, offering the practical advantage that less culture medium changes are required. However, these results should be interpreted with caution and further reproduction is required before possible biological explanations are discussed.
Due to the non-linearity of the 4pLL model, the function was approximated according to the least square method with the Gauss–Newton algorithm.
After cell counting using Trypan blue to determine viability, 50,000 cells in FCS-containing medium were plated into each well of 96-well plates and kept at 37 °C for at least 3 h.
For single exposure, the cells were incubated with compounds for 24 h or 48 h; for repeated exposure, the compound-containing medium was renewed every 48 h and the cells were incubated for a total of 7 days.
washout experiments with an initial exposure period with test compounds followed by test compound-free incubation or repeated exposure and washout periods may be considered.
The advantage of this fitting procedure is that the left asymptote is used as a control level for calculation of EC50 and EC20 values which are more robust than just using the values of the solvent controls.
Only when the cytotoxic test compound concentrations were not reached with 0.1% DMSO, the solvent concentration was increased to 0.5%.
Compounds with a relatively strong decrease in EC50 after 7 days compared to 1 day are busulfan, famotidine, and isoniazid (Fig. 2b).
The applied solvent concentrations of 0.1% or 0.5% DMSO did not cause any cytotoxicity compared to cells cultivated in medium without DMSO.
Dimethyl sulfoxide DILI Drug-induced liver injury EtOH Ethanol FAM Famotidine Glc Glucose HYZ Hydroxyzine INAH Isoniazid KC Ketoconazole LAB Labetalol LEV Levofloxacin MEL Melatonin MePa Methylparaben NAC N-Acetylcysteine NIM Nimesulide NFT Nitrofurantoin PhB Phenylbutazone PMZ Promethazine PPL Propranolol RIF Rifampicin TSN Triclosan VPA Valproic acid Vit C Vitamin CDrug-induced liver injury (DILI) is one of the principal reasons for drug withdrawal from the market (Godoy et al. 2013; Hewitt et al. 2007).
large efforts are undertaken to establish in vitro tests with the long-term goal to predict human toxicity (Daneshian et al.