A Comparison between a Manual and Automatic Method to
Characterize Red Blood Cell Trajectories
Diana Pinho*, Fernando Gayubo
§
, Ana I. Pereira*
,£
and Rui Lima*
,
**
*
Polytechnic Institute of Bragança, Portugal
§
Fundación CARTIF, Valladolid, Spain
£
Algoritmi, University of Minho, Portugal
**
CEFT, FEUP, Porto University, Portugal
Abstract. The current study proposes an automatic method for the segmentation and tracking of red blood cells (RBCs)
flowing through a 100 m glass capillary. The measurements obtained with the proposed automatic method are compared
with a manual tracking method using nonlinear optimization techniques.
Keywords: Automatic Method. Red Blood Cells. Nonlinear Optimization.
PACS:
02.60.Pn
INTRODUCTION
Blood flow in microcirculation is crucial for the normal function of tissues and organs. Detailed study of blood
cells flowing in microvessels and microchannels is essential to provide a better understanding on the blood
rheological properties and disorders in microcirculation [1-5]. Image analysis plays an extremely important role to
obtain information about the blood rheology. However, most of the past blood flow experimental data have been
performed manually [1-3]. Although manual methods can be highly reliable, these methods are relatively time
consuming and can also introduce user errors into the data. As a result, it is crucial to develop image analysis
methods able to get the data automatically. The present paper presents an approach able to track the RBCs
trajectories automatically. Nonlinear optimization techniques were used to compare automatic and manual methods.
MATERIALS AND METHODS
Experimental Set-up
The confocal system used in this study consists of an inverted microscope (IX71; Olympus) combined with a
confocal scanning unit (CSU22; Yokogawa), a diode-pumped solid-state (DPSS) laser (Laser Quantum) with an
excitation wavelength of 532 nm and a high-speed camera (Phantom v7.1; Vision Research). The laser beam was
illuminated from below the microscope stage through a dry 40u objective lens. The glass capillary was placed on the
stage of the inverted microscope and by using a syringe pump (KD Scientific) a pressure-driven flow was kept
constant (Re ~ 0.008). More detailed information about this system can be found elsewhere [1].
The confocal images were captured around the middle of the capillary with a resolution of 640u480 pixel at a
rate of 100 frames/s. Two image analyses methods were used in this study: manual method (MM) and automatic
method (AM).
Numerical Analysis and Applied Mathematics ICNAAM 2011
AIP Conf. Proc. 1389, 767-770 (2011); doi: 10.1063/1.3636845
© 2011 American Institute of Physics 978-0-7354-0956-9/$30.00
767

A man
u
individual
through su
c
for the det
e
All fra
m
images wit
h
most of th
e
next step,
t
RBCs fro
m
image). Th
to divide t
h
images.
The obj
e
computes
a
Sobel oper
a
about this
a
u
al tracking
p
RBC. By usi
n
c
cessive ima
g
e
rmination of
m
es were loa
d
h
the functio
n
e
noise and to
t
he images a
r
m
the backgro
u
is is possible
h
e image into
o
FIGURE 1. T
e
cts are defin
e
a
n approxima
t
ator is either
a
utomatic me
t
p
lugin (MTra
c
n
g MTrackJ
p
g
es for an inte
r
each individ
u
d
ed and pre-
p
n
imcrop. Th
e
enhance the
f
r
e subject to
a
u
nd, i.e. diffe
r
using a thres
h
o
ne or more r
he region of in
t
e
d with the S
o
t
ion of the gr
a
the correspo
t
ho
d
can be f
o
FIGURE 2.
R
Ma
n
c
kJ) [6] of a
n
p
lugin, the b
r
r
val of time o
al RBC traje
c
A
uto
m
p
rocessed usi
n
e
median func
t
f
lowing obje
c
a
segmentati
o
r
entiate the a
r
h
old method,
egions. The f
u
t
erest (above)
a
o
bel filter (s
e
a
dient of the i
m
n
ding gradie
n
o
und elsewhe
r
R
esult of the it
e
n
ual Method
(
n
image anal
y
r
ight centroi
d
f 10 ms. Afte
r
c
tory.
m
atic Metho
d
n
g Matlab [8]
tion, medfilt2
c
t. In Fig. 1
w
o
n filter, Sob
e
r
ea of interes
t
where a defi
n
u
nction iterat
i
a
nd the image f
i
e
e Fig. 2), w
h
mage intensi
t
n
t vector or
t
r
e [9].
e
rative threshol
d
(
MM)
y
sis software
d
of the selec
t
r
obtaining x
a
d
(AM)
. The region
, with one m
a
w
e can see the
e
l. With this
s
t
(the RBCs)
f
n
ition of one
o
i
ve threshold
w
i
ltered by usin
g
h
ich shows o
n
t
y. At each pi
x
t
he norm of
t
d
method and t
h
(Image J, NI
H
t
ed RBC was
a
nd y
p
ositio
n
of interest w
a
a
sk 5x5 pixel,
result of thes
s
egmentation
f
rom the no
t
-
i
o
r more valu
e
w
as applied f
o
g
the median fu
n
n
ly the edge
o
x
el point in t
h
h
is vec
t
or.
M
h
e filter Sobel.
H
) [7] was u
s
automatical
l
n
s, the data w
e
a
s then crop
p
,
was applied
s
e processing
it is possibl
e
interest area
(
e
s of separati
o
fo
r the sequen
c
n
ction medfilt2
o
f the object
s
h
e image, the
M
ore detailed
sed to track
y computed
e
re exported
p
ed from the
to eliminate
steps. In the
e
to separate
(
background
o
n is enough
c
e of all the
.
s
. The Sobel
result of the
information
768

RESULTS AND DISCUSSION
After obtaining series of ݔ and ݕ positions for both methods, data were exported for the determination of each
individual RBC trajectory and to analyze the best mathematical function that approximates to the RBCs
experimental flow behavior.
In this study it was measured and analyzed twenty five red blood cells at a temperature of 25 ºC. For each cell ݅,
by using the MTrackJ plugin (MM) form ImageJ and the propose method (AM), we have obtained ൛൫ݔ
ǡݕ
൯ǡ ݆ൌ
ͳǡǥǡ݇
ൟ data. To compare the consistency of both methods we decided to determine the better approximation for
the data by using a nonlinear optimization [10]. Hence, we consider three different functions [5] (polynomials, sum
of
trigonometric functions and a sum of exponential functions) defined as:
݂
ଵ
ሺ
ݔǡ
ሻ
ൌ
ݔ
ଽ
ୀ
Ǣ݂
ଶ
ሺ
ݔǡܽǡܾ
ሻ
ൌܽ
ܽ
଼
ୀଵ
ሺሻܾ
ሺ
݅ݔ
ሻ
Ǣ݂
ଷ
ሺ
ݔǡܿǡ݀ǡ݃
ሻ
ൌ݃
݁
ቆ
ି
൬
ሺ
௫ି
ሻ
ௗ
൰
మ
ቇ
଼
ୀଵ
Ǥ
where אܫܴ
ͳͲ
, ܽאܫܴ
ͻ
ǡܾǡܿǡ݀ǡ݃אܫܴ
଼
are the function parameters and the vector ݔאܫܴ
݇݅
, where ݅
represents the cell number. To identify the functions parameters it was used the tool cftool present in Curve
Fitting Toolbox from Matlab [11].
The error of nonlinear least squares approximation of the selected RBCs are listed in the Table 1, where RBC
refers to the red blood cell number, ܨ
ൌ
σ
൫ݕ
െ݂
ሺ
ݔ
ሻ
൯
ଶ
ୀ
is the nonlinear least squares approximation error of the
function ݂
ሺݔǡǤሻǡ with ݅ൌͳǡʹǡ͵, Av refers the error average, and s corresponds to the standard deviation of the
errors.
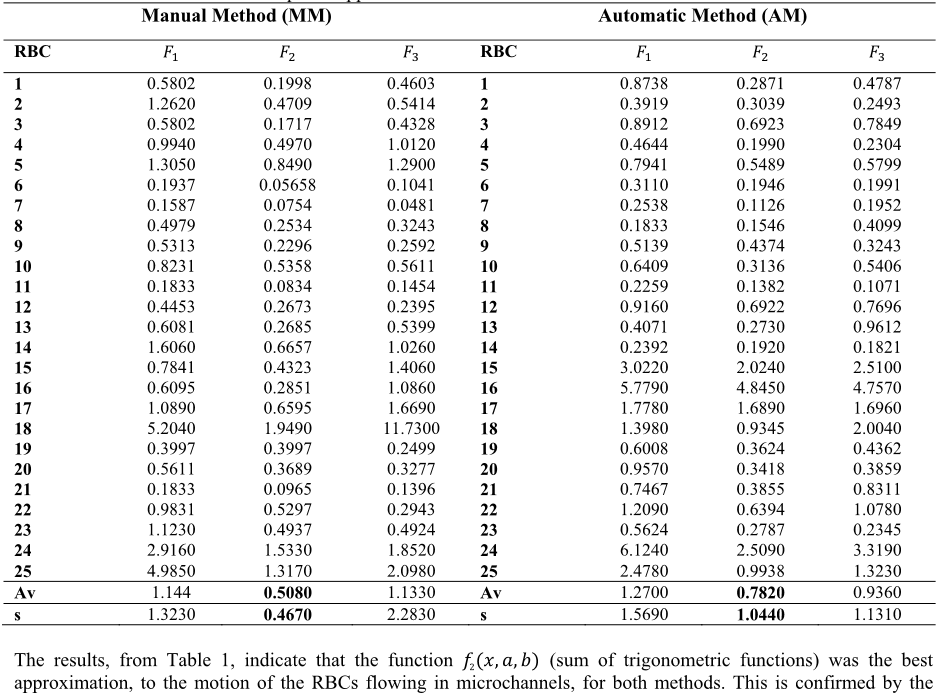
Table 1 - Errors of nonlinear least squares approximation of the selected RBCs for both methods.
Manual Method (MM) Automatic Method (AM)
RBC
ܨ
ଵ
ܨ
ଶ
ܨ
ଷ
RBC
ܨ
ଵ
ܨ
ଶ
ܨ
ଷ
1
0.5802 0.1998 0.4603
1
0.8738 0.2871 0.4787
2
1.2620 0.4709 0.5414
2
0.3919 0.3039 0.2493
3
0.5802 0.1717 0.4328
3
0.8912 0.6923 0.7849
4
0.9940 0.4970 1.0120
4
0.4644 0.1990 0.2304
5
1.3050 0.8490 1.2900
5
0.7941 0.5489 0.5799
6
0.1937 0.05658 0.1041
6
0.3110 0.1946 0.1991
7
0.1587 0.0754 0.0481
7
0.2538 0.1126 0.1952
8
0.4979 0.2534 0.3243
8
0.1833 0.1546 0.4099
9
0.5313 0.2296 0.2592
9
0.5139 0.4374 0.3243
10
0.8231 0.5358 0.5611
10
0.6409 0.3136 0.5406
11
0.1833 0.0834 0.1454
11
0.2259 0.1382 0.1071
12
0.4453 0.2673 0.2395
12
0.9160 0.6922 0.7696
13
0.6081 0.2685 0.5399
13
0.4071 0.2730 0.9612
14
1.6060 0.6657 1.0260
14
0.2392 0.1920 0.1821
15
0.7841 0.4323 1.4060
15
3.0220 2.0240 2.5100
16
0.6095 0.2851 1.0860
16
5.7790 4.8450 4.7570
17
1.0890 0.6595 1.6690
17
1.7780 1.6890 1.6960
18
5.2040 1.9490 11.7300
18
1.3980 0.9345 2.0040
19
0.3997 0.3997 0.2499
19
0.6008 0.3624 0.4362
20
0.5611 0.3689 0.3277
20
0.9570 0.3418 0.3859
21
0.1833 0.0965 0.1396
21
0.7467 0.3855 0.8311
22
0.9831 0.5297 0.2943
22
1.2090 0.6394 1.0780
23
1.1230 0.4937 0.4924
23
0.5624 0.2787 0.2345
24
2.9160 1.5330 1.8520
24
6.1240 2.5090 3.3190
25
4.9850 1.3170 2.0980
25
2.4780 0.9938 1.3230
Av
1.144
0.5080
1.1330
Av
1.2700
0.7820
0.9360
s
1.3230
0.4670
2.2830
s
1.5690
1.0440
1.1310
The results, from Table 1, indicate that the function ݂
ʹ
ሺݔǡܽǡܾሻ (sum of trigonometric functions) was the best
approximation, to the motion of the RBCs flowing in microchannels, for both methods. This is confirmed by the
769

value of the error average. Another important aspect is the fact that the standard deviation of the errors is small when
we use the function ݂
ʹ
.
CONCLUSIONS
The present study indicates that the data obtained from the proposed automatic method (AM) is equivalent to
data obtained from the manual method (MM) as the function that best approaches is the same, i.e., the function ݂
ଶ
(sum of trigonometric functions). Hence, the proposed automatic method is a promising way to track blood cells
flowing in microchannels.
ACKNOWLEDGMENTS
The authors acknowledge the financial support provided by: PTDC/SAU-BEB/108728/2008, PTDC/SAU-
BEB/105650/2008 and PTDC/EME-MFE/099109/2008 from the FCT (Science and Technology Foundation) and
COMPETE, Portugal.
REFERENCES
1. R. Lima, T. Ishikawa, Y. Imai, M. Takeda, S. Wada, and T. Yamaguchi, Measurement of individual red blood
cell motions under high hematocrit conditions using a confocal micro-PTV system. Annals of Biomedical
Engineering. 37, 1546-59, 2009.
2. H. Fujiwara, T. Ishikawa, R. Lima, et al. Red blood cell motions in high-hematocrit blood flowing through a
stenosed microchannel. Journal of Biomechanics 42, 838-843, 2009.
3. Y. Suzuki, N. Tateishi, M. Soutani and N. Maeda, Deformation of erythrocytes in microvessels and glass
capillaries: effects of erythrocyte deformability. Microcirculation 3, 49-57, 1996.
4. A. Pries, T. Secomb, et al, Resistance to blood flow in microvessels in vivo. Circulation Research 75, 904-915,
1994.
5. D. Pinho, et al., Red blood cells motion in a glass microchannel, Numerical Analysis and Applied Mathematics,
Vol. 1281: 963-966, 2010.
6. E. Meijering, I. Smal and G. Danuser, Tracking in molecular bioimaging, IEEE Signal Process. Mag. 23: 46–53,
2006.
7. M. Abramoff, P. Magelhaes and S. Ram, Image processing with image J, Biophotonics Int. 11: 36–42, 2004.
8. S. L. Eddins, R. C. Gonzalez, R. E. Woods, Digital Image Processing Using Matlab, 2002.
9. D. Pinho, Determination and characterization of red blood cells trajectories: a semi-automatic method, Master in
Biomedical Technology, Polytechnic Institute of Bragança, Portugal, 2011 (in portuguese).
10. J. Nocedal and S. Wright, Numerical Optimization. Springer Series in Operations Research, Springer, New
York, 1999.
11. MathWorks, Optimization Toolbox User’s Guide, 2010.
770