Acinetobacter nectaris sp. nov. and Acinetobacter
boissieri sp. nov., isolated from floral nectar of wild
Mediterranean insect-pollinated plants
Sergio A
´
lvarez-Pe
´
rez,
1
3 Bart Lievens,
2,3
3 Hans Jacquemyn
4
and Carlos M. Herrera
1
Correspondence
S. A
´
lvarez-Pe
´
rez
sealperez@gmail.com
B. Lievens
bli@scientiaterrae.org
1
Estacio
´
n Biolo
´
gica de Don
˜
ana, Consejo Superior de Investigaciones Cientı´ficas (CSIC), Avda.
Ame
´
rico Vespucio, E-41092 Sevilla, Spain
2
Laboratory for Process Microbial Ecology and Bioinspirational Management (PME&BIM), Lessius
University College, De Nayer Campus, Consortium for Industrial Microbiology and Biotechnology
(CIMB), Department of Microbial and Molecular Systems (M
2
S), KU Leuven Association, B-2860
Sint-Katelijne-Waver, Belgium
3
Scientia Terrae Research Institute, B-2860 Sint-Katelijne-Waver, Belgium
4
Division of Plant Ecology and Systematics, Biology Department, KU Leuven, B-3001 Heverlee,
Belgium
The taxonomic status of 14 strains of members of the genus Acinetobacter isolated from floral
nectar of wild Mediterranean insect-pollinated plants, which did not belong to any previously
described species within this genus, was investigated following a polyphasic approach.
Confirmation that these strains formed two separate lineages within the genus Acinetobacter was
obtained from comparative analysis of the partial sequences of the 16S rRNA gene and the gene
encoding the b-subunit of RNA polymerase ( rpoB), DNA–DNA reassociation data, determination
of the DNA G +C content and physiological tests. The names Acinetobacter nectaris sp. nov. and
Acinetobacter boissieri sp. nov. are proposed. The type strain of A. nectaris sp. nov. is SAP
763.2
T
(5LMG 26958
T
5CECT 8127
T
) and that of A. boissieri sp. nov. is SAP 284.1
T
(5LMG 26959
T
5CECT 8128
T
).
Members of the genus Acinetobacter are generally regarded
as common, free-living saprophytes that show extensive
metabolic versatility and potential to adapt to different
human-associated and natural environments (Towner,
2006; Doughari et al. , 2011; Sand et al., 2011). Apart from
the well-known human and animal-pathogenic species of
the genus Acinetobacter , several novel species within this
genus have been described during recent years to
accommodate isolates from agricultural soils (Kang et al. ,
2011), activated sludge (Carr et al. , 2003), raw wastewater
(Vaz-Moreira et al., 2011), a hexachlorocyclohexane
dumpsite (Malhotra et al. , 2012) and diverse natural
environmental sources, such as forest soils (Kim et al.,
2008), seawater (Vaneechoutte et al., 2009) and wetlands
(Anandham et al. , 2010). Nevertheless, except for those
species with clinical importance, the distribution and
ecological role(s) of the ‘acinetobacters’ in most environ-
ments are largely unknown (Carr et al., 2003; Towner,
2006). In particular, the possible associations of members
of this bacterial group with plant hosts remain to be
addressed.
Floral nectar is the key component in the mutualism
between angiosperms and their animal pollinators, which
take this sugary solution as a reward for their pollination
services (Brandenburg et al., 2009; Heil, 2011). While
foraging on flowers, pollinators can contaminate floral
nectar with different prokaryotic and eukaryotic micro-
organisms, some of which are particularly well-adapted to
thrive in this ephemeral habitat characterized by high
osmotic pressure and the presence of plant secondary
metabolites with defensive functions (Herrera et al. , 2010;
Pozo et al. , 2012). Nectar micro-organisms can alter
pollinators’ foraging behaviour in different ways, for
example by reducing the nutritional value of floral nectar
3These authors contributed equally to this work.
Abbreviations:
BI, Bayesian inference; ML, maximum-likelihood; NJ,
neighbour-joining; OTU, operational taxonomical unit; PM, Phenotype
MicroArray.
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene
sequences determined in this study are JQ771129–JQ771142 and
those for the partial rpoB gene sequences are JQ771143–JQ771156.
A supplementary figure and four supplementary tables are available with
the online version of this paper.

and/or changing other physico-chemical conditions within
the floral microenvironment, and thus potentially interfere
with plant sexual reproduction, as has been recently
suggested for nectar yeasts (Canto et al. , 2008; Herrera
et al., 2008; de Vega et al. , 2009; Herrera & Pozo, 2010;
Peay et al., 2012).
Recently, species of the genus Acinetobacter have been
identified as representing the main bacterial genus
inhabiting the floral nectar of some cultivated plant species
from Northern Israel (Fridman et al. , 2012) and phylo-
genetically diverse wild Mediterranean plants from
Southern Spain (A
´
lvarez-Pe
´
rez & Herrera, 2013). In this
latter study, nectar isolates of members of the genus
Acinetobacter grouped into a single operational taxonom-
ical unit (OTU) defined on the basis of a 3 % dissimilarity
cut-off in the 16S rRNA gene sequence, but into two
different OTUs when this threshold was lowered to 1 %
(A
´
lvarez-Pe
´
rez & Herrera, 2013). Such differentiation was
also supported by some differences in colony morphology
and growth rate in plate cultures, but no additional tests
were conducted to further characterize those isolates. In
this study we explore the taxonomic status of these two
nectar groups of acinetobacters associated with wild
Mediterranean plants.
The 14 strains investigated in this study are listed in Table
S1, available in IJSEM Online. These strains were isolated
on different dates from nectar samples of several plant
species collected at different places within the surroundings
of Don
˜
ana’s Natural Park (Huelva province, southwest
Spain), using the procedure described by A
´
lvarez-Pe
´
rez
et al. (2012). Additionally, for comparative taxonomic
analysis, Acinetobacter calcoaceticus DSM 30006
T
,
Acinetobacter baylyi DSM 14961
T
, Acinetobacter gerneri
DSM 14967
T
and Acinetobacter radioresistens DSM 6976
T
were included in some phenotypic and genotypic assays.
The inclusion of Acinetobacter calcoaceticus in those tests
was justified by its status as the type species for the genus
Acinetobacter, while the other three taxa were identified as
the most closely related species to our nectar strains on the
basis of 16S rRNA gene sequence data (see below).
Colonies of all the nectar strains grown on trypticase soy
agar (TSA; Panreac) were circular, convex to umbilicate,
smooth and slightly opaque with entire margins. After
5 days of incubation at 25
u
C, colonies of these strains were
variable in diameter, ranging from 0.5 to 2.5 mm, although
smaller colonies were also observed for some strains. On
microscope images, cells appeared as non-motile coccoba-
cilli and commonly occurred in pairs, but also alone or in
short chains. No spores were observed. The main
phenotypic properties of the type strains of all studied
taxa are summarized in Table 1, and those of all studied
nectar isolates are shown in Table S2. All tests were carried
out at 25
u
C unless otherwise indicated. Catalase activity
was determined by evaluating bubble production with a
3 % (v/v) hydrogen peroxide solution (Cappuccino &
Sherman, 2002). Oxidase activity was tested using oxidase
test strips (MB 0266 A; Oxoid,). All strains were aerobic,
catalase-positive and oxidase-negative. Whereas no growth
was observed on TSA in an anaerobic jar, all nectar strains
were able to grow at decreased oxygen levels, as assessed by
visual inspection of bacterial cultures grown on TSA in a
candle jar for 5 days at 25
u
C. Growth at 4, 25, 30, 37 and
41
u
C, haemolysis of sheep blood, gelatin hydrolysis and
production of acid from glucose and sucrose in Hugh–
Leifson medium were examined as described previously
(Hugh & Leifson, 1953; Skerman, 1959; Bouvet & Grimont,
1986; Carr et al., 2003). All nectar strains grew at 25 and
30
u
C, but not at 37 or 41
u
C. Strains SAP219.2, SAP239.2,
SAP240.2, SAP241.2, SAP242.2, SAP284.1
T
and SAP320.1
were able to grow at 4
u
C but strains SAP220.2, SAP249.1,
SAP 305.1, SAP763.2
T
, SAP 956.2, SAP970.1 and SAP971.1
could not. All strains were non-haemolytic on Columbia
agar supplemented with sheep blood. Strains SAP 305.1
and SAP 970.1 were found to grow poorly on this medium.
None of the tested isolates were able to hydrolyse gelatin.
All nectar strains produced acid from sucrose and glucose.
Carbon source oxidation was determined by Phenotype
MicroArray (PM) technology (Biolog) using PM plate 1.
Using this technology, kinetic profiles are generated by
continuously monitoring the metabolic activity during
incubation (Bochner et al., 2001). Plates were incubated in
the OmniLog automated incubator-reader (Biolog) for
5 days at 25
u
C and were read every 15 min. Interpretation
of results was performed using OmniLog PM software
according to the manufacturer’s instructions. Clear differ-
ences were observed between the two groups of nectar
strains and between these nectar strains and the type strains
of A. baylyi , A. calcoaceticus , A. gerneri and A. radioresistens .
In contrast to the type strains of the related species of the
genus Acinetobacter , all nectar strains were able to oxidize
sucrose and
D-fructose as the only carbon source. D-
mannose, on the other hand, was only oxidized by the
group which included strain SAP763.2
T
. In addition, D-
glucose was only oxidized by some strains of that group. In
addition to sucrose and
D-fructose, L-malic acid was the
only carbon source that could be oxidized by the majority
of the strains of the group containing strain SAP284.1
T
. For
the group including strain SAP763.2
T
, all strains were
found to oxidize
L-aspartic acid, bromosuccinic acid,
fumaric acid,
L-glutamic acid, L-malic acid, DL-malic acid,
succinic acid,
L-asparagine and L-proline; and some of the
seven strains in this group were able to oxidize
D-xylose, D-
gluconic acid,
a-ketoglutaric acid, mono-methylsuccinate
or
L-alanine (Table S2). In order to compare the results
obtained by PM fingerprinting with more conventional
assimilation tests commonly used for classification of
members of the genus Acinetobacter , some key biochemical
features were also assessed using the phenotypic system
described by Bouvet & Grimont (1986) and adapted by
Nemec et al. (2009). More specifically, tests were
performed for sucrose,
D-glucose, succinic acid and
phenylacetate. Briefly, assimilation tests were performed
using the basal mineral medium of Cruze et al. (1979)

supplemented with 0.1 % (w/v) of the tested carbon source.
The basal medium consisted of the following (l
21
): 10.0 g
KH
2
PO
4
,5.0gNa
2
HPO
4
, 2.0 g (NH
4
)
2
SO
4
, 0.2 g
MgSO
4
.7H
2
O, 0.001 g CaCl
2
.2H
2
O and 0.001 g
FeSO
4
.7H
2
O (pH 7.0). 5 ml of this medium was dispensed
into tubes, inoculated with washed bacterial cells and
incubated at 25
u
C under agitation. Growth on the different
carbon sources was evaluated after 2, 4, 6 and 10 days by
means of visual comparison between inoculated tubes
containing carbon sources and control tubes containing
only inoculated basal medium. In general, results obtained
by these assays confirmed the results obtained by the PM
technology assays (data not shown). From the two groups of
nectar isolates, all isolates were again able to assimilate
sucrose, whereas
D-glucose and succinic acid were only
assimilated by (some of) the isolates of the group which
included SAP763.2
T
. In contrast to other known species of
the genus Acinetobacter , none of the nectar isolates were able
to assimilate acetate.
Sucrose tolerance was determined by culturing the studied
strains in transparent plastic vials containing Luria–Bertani
(LB) broth (Difco) supplemented with 0 (positive control),
10, 20, 30, 40 or 50 % sucrose (w/v, Sigma–Aldrich). All
these liquid media were filter-sterilized and kept at 4
u
C
until use. The range of sugar concentrations tested closely
Table 1. Differential phenotypic characteristics of the type
strains of all studied species of the genus Acinetobacter
Strains: 1, A. nectaris sp. nov. SAP 763.2
T
;2,A. boissieri sp. nov. SAP
284.1
T
;3,A. calcoaceticus DSM 30006
T
;4,A. baylyi DSM 14961
T
;5,A.
gerneri DSM 14967
T
;6,A. radioresistens DSM 6976
T
. A. nectaris sp.
nov. and A. boissieri sp. nov. can be separated from the type strains of
A. calcoaceticus , A. baylyi , A. gerneri and A. radioresistens by some
basic phenotypic characteristics, such as their ability to oxidize
fructose and sucrose, their tolerance to sucrose concentrations above
30 % (w/v) and their inability to grow at ¢37
u
C and assimilate
acetate. + , Positive reaction; 2, negative reaction;
W, weak growth;
ND, no data available.
Characteristic 1 2 3 4 5 6
Growth on TSA at:
4
u
C 2 +
WW2 +
25
u
C ++ + + + +
30
u
C ++ + + + +
37
u
C 22 ++++
41
u
C 22 2
W ++
Anaerobic growth 222222
Growth at decreased oxygen levels ++
ND ND ND ND
Haemolysis on Columbia blood agar 222222
Gelatin hydrolysis 222222
Catalase activity ++ + + + +
Oxidase activity 222222
Growth on LB broth plus sucrose at:
10 % (w/v) ++ + + + +
20 % (w/v) ++ + + + +
30 % (w/v) ++ + + + 2
40 % (w/v) ++ 2222
50 % (w/v) +
W 2222
Acid production from glucose ++ + + + 2
Acid production from sucrose ++ + + + 2
Oxidation of: *
D-Glucose 22 2 + 22
D-Fructose ++ 2222
D-Mannose + 22222
Sucrose ++ 2222
D-Xylose 222222
Acetic acid 22 ++++
D-Aspartic acid 22 2 + 22
L-Aspartic acid + 22 + 22
Bromosuccinic acid + 22 ++2
Citric acid 22 2 ++2
Fumaric acid + 22 ++2
D-Galacturonic acid 22 + 222
D-Gluconic acid + 22 + 22
L-Glutamic acid + 22 ++2
a-Hydroxybutyric acid 22 2 ++2
m-Hydroxyphenyl acetic acid 22 2 2 + 2
p-Hydroxyphenyl acetic acid
22 2 2
+
2
a-Ketobutyric acid 22 2 ++2
a-Ketoglutaric acid 22 2 ++2
L-Lactic acid 22 2 ++2
D-Malic acid
222222
L-Malic acid ++ 2 ++2
DL-Malic acid + 22 ++2
Table 1. cont.
Characteristic 1 2 3 4 5 6
Mucic acid 22 2 + 22
Propionic acid 22 2 ++2
Pyruvic acid 22 2 ++2
D-Saccharic acid 22 2 + 22
Succinic acid + 2 +++2
Tricarballylic acid 22 2 + 22
Methylpyruvate 22 +++2
Mono-methylsuccinate 22 2 ++2
a-Hydroxyglutaric acid-g-Lactone 22 2 2 + 2
2-Aminoethanol 222222
Dulcitol
22
+
222
D-Alanine 22 +++2
L-Alanine 22 2 ++2
L-Asparagine + 22 + 22
L-Glutamine + 22 ++2
Gly–Pro 22 2 2 + 2
Phenylethylamine 22 2 2 + 2
L-Proline + 22 ++2
L-Threonine 22 + 222
Tween 20 22 +++2
Tween 40 22 +++2
Tween 80 22 ++++
*Oxidation of carbon sources was determined by Phenotype
MicroArray (PM) technology (Biolog) using PM plate 1. Further
details on the procedure and the results obtained for all the novel
nectar isolates characterized in this work are provided in Table S2.

matched the range of naturally occurring variation in floral
nectars of the wild Mediterranean plant communities from
which our nectar strains were recovered (S. A
´
lvarez-Pe
´
rez
& C. M. Herrera, unpublished results). Single colonies
picked from 5-day cultures on TSA medium were used to
inoculate the tubes and these were incubated at 25
u
C for
up to 10 days. The turbidity of the cultures with respect to
negative controls (i.e. tubes containing no inoculated
media) was recorded as a positive result. At the end of the
experiment, an aliquot of each test tube was plated on
TSA medium to check for possible contaminations.
Furthermore, acid production by bacterial strains when
growing at different sucrose concentrations was tested by
adding 40
ml methyl red (Panreac) to each tube. All nectar
strains were able to grow at sucrose concentrations ranging
from 10 to 50 % (w/v), although the growth of strains of
the group containing SAP284.1
T
at 40 % and 50 % sucrose
was very weak. Acidification of the culture media was
observed in all tubes containing sucrose, but not in the
positive control (no sucrose). In contrast, A. baylyi DSM
14961
T
, A. calcoaceticus DSM 30006
T
and A. gerneri DSM
14967
T
only grew at sucrose concentrations up to 30 %,
and A. radioresistens DSM 6976
T
only tolerated 10 % and
20 % sucrose. Furthermore, these four reference strains did
not acidify either sucrose-containing culture broths or the
positive control.
Methods for genotypic characterization of the studied
strains included comparative sequence analysis of the 16S
rRNA and the
b-subunit of RNA polymerase (rpoB)-
encoding genes, assessment of overall genomic relatedness
by DNA–DNA hybridizations and determination of the
DNA G+C content.
An almost complete fragment of the 16S rRNA gene
was amplified and subsequently sequenced as described
by A
´
lvarez-Pe
´
rez et al. (2012). Preliminary sequence
comparisons with the 16S rRNA gene sequences stored in
GenBank (http://www.ncbi.nlm.nih.gov/genbank/) and the
Ribosomal Database Project (RDP, http://rdp.cme.msu.
edu/) databases showed that the nectar strains belonged to
the family Moraxellaceae in the Gammaproteobacteria
subdivision, and the best hits for all sequences were the
putative isolates of members of the genus Acinetobacter
recovered by Fridman et al. (2012) from floral nectar of
cultivated plants (99 % and 97–98 % overall similarity to
strains SAP763.2
T
and SAP284.1
T
, respectively). The
SimTable tool available at the EzTaxon server v. 2.1
(http://www.eztaxon.org/, last accessed 10 December 2011;
Chun et al. , 2007) was used to search for neighbours
among species of the genus Acinetobacter with validly
published names on the basis of 16S rRNA gene sequences,
identifying A. baylyi B2
T
, A. gerneri 9A01
T
and A.
radioresistens DSM 6976
T
as the species most closely
related to the novel nectar strains, but with a sequence
similarity value ¡ 96.3 % in all cases (Table S3). The
sequence similarity value between strains SAP 763.2
T
and
SAP 284.1
T
, as determined through the EzTaxon server,
was 97.7 %.
The 16S rRNA gene sequences of the novel nectar strains
and reference strains of members of the genus Acinetobacter
and the family Moraxellaceae were included in a multiple
alignment generated by
CLUSTAL W (Chenna et al., 2003).
The resulting alignment was trimmed with BioEdit v.
7.0.9.0 (Hall, 1999) to ensure that all sequences had the
same start and end point, and analysed with Gblocks
(Castresana, 2000) to eliminate ambiguously aligned
regions, using ‘allow gap positions5with half’, ‘minimum
length of a block
5
5
9
and default settings for all other
options. Following these procedures, 1320 nt positions
(98 % of the original alignment) remained for subsequent
phylogenetic analysis using the neighbour-joining (NJ)
method as implemented in the
MEGA 5 software package
(Tamura et al. , 2011). Pairwise evolutionary distances were
computed by the Jukes–Cantor method, and reliability of
nodes in the NJ phylogram was assessed by running 1000
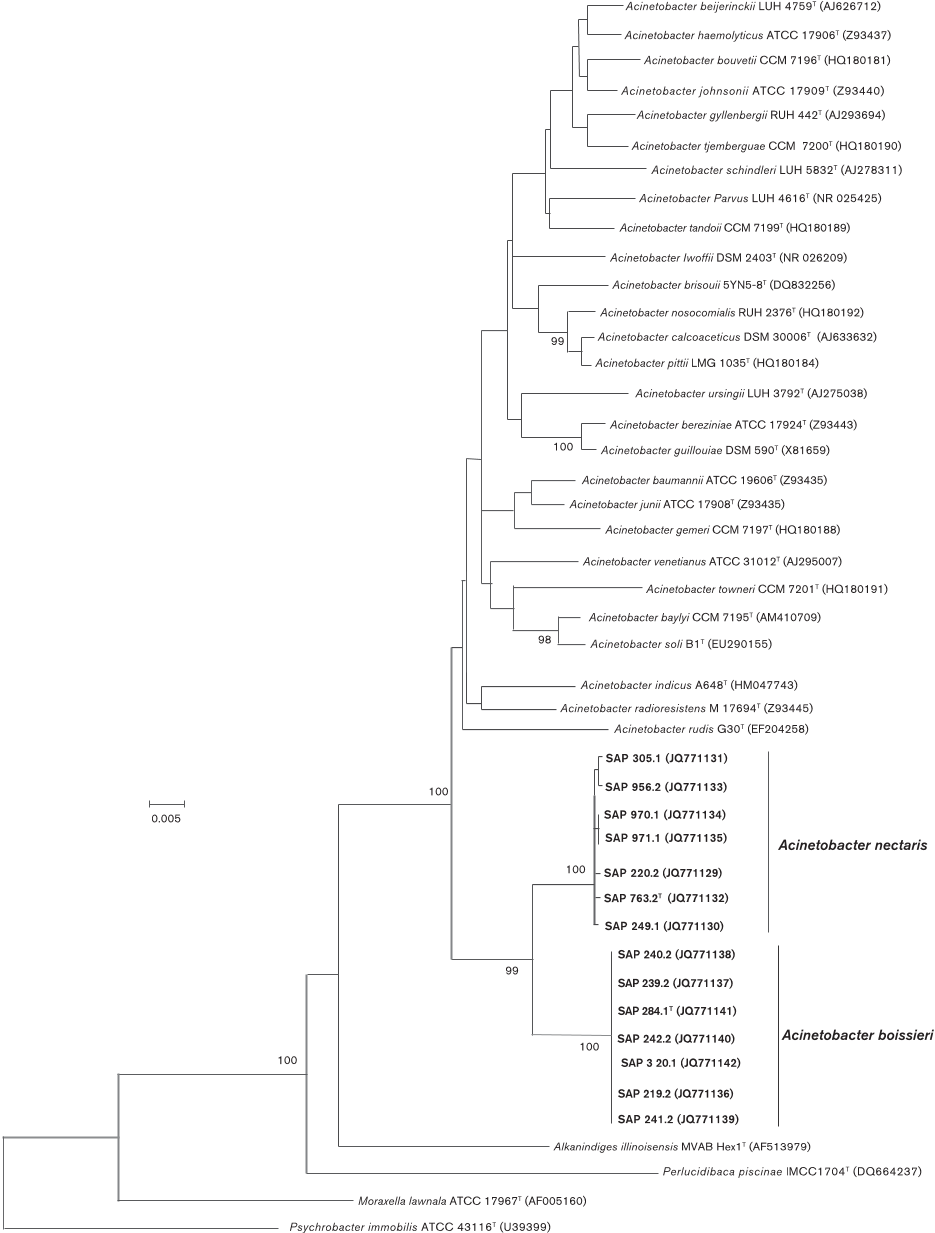
bootstrap replicates. In the NJ phylogram based on 16S
rRNA gene sequences (Fig. 1) the novel nectar strains
clustered with other members of the genus Acinetobacter ,
but stood apart from the recognised species of this genus
by forming a consistently differentiated group with 99 %
bootstrap support. Furthermore, all strains of the group
which included strain SAP 763.2
T
clustered together with a
100 % bootstrap support, as did all those strains corres-
ponding to the group which included strain SAP 284.1
T
(Fig. 1).
Comparative sequence analysis of two variable regions –
zone 1 (approximately 397 bp) and zone 2 (approximately
544 bp) – of the rpoB gene was used to confirm both the
within-species relatedness of the two groups of novel
strains, and their separation from each other and from
previously described species of the genus Acinetobacter .
Primer sequences and PCR conditions were as previously
described (Khamis et al. , 2004; La Scola et al., 2006), with
some minor modifications: 250
mM of each dNTP (Sigma–
Aldrich), 0.4
mM of each of the corresponding forward and
reverse primers (Sigma–Aldrich) and 5610
22
U Taq
polymerase
ml
21
(Bioline) were used in reaction mixtures
and 52
u
C was the temperature for primer annealing in
PCR cycles. As with the 16S rRNA gene sequence,
concatenated sequences of rpoB zones 1 and 2 of the
nectar strains and type strains of other species of the genus
Acinetobacter were included in a multiple alignment and
analysed with Gblocks, which resulted in the selection of
843 nucleotide positions (99 % of the original alignment).
A phylogenetic tree was inferred by the NJ method using
the Jukes–Cantor method, and again, the two groups of
nectar strains formed two different clusters with bootstrap
values supporting their distinctness from each other and
from the other acinetobacters (Fig. S1).
The genomic DNA–DNA relatedness between the strains
SAP 763.2
T
and SAP 284.1
T
and between these and the type
strains of A. calcoaceticus , A. baylyi and A. gerneri was
evaluated by DNA–DNA hybridizations. High-molecular-
mass total genomic DNA was extracted by the method of
Wilson (1987). DNA–DNA hybridizations were carried out
0.005
Acinetobacter beijerinckii
LUH 4759
T
(AJ626712)
Acinetobacter haemolyticus
ATCC 17906
T
(Z93437)
Acinetobacter bouvetii
CCM 7196
T
(HQ180181)
Acinetobacter johnsonii ATCC 17909
T
(Z93440)
Acinetobacter gyllenbergii
RUH 442
T
(AJ293694)
Acinetobacter tjemberguae
CCM 7200
T
(HQ180190)
Acinetobacter schindleri LUH 5832
T
(AJ278311)
.
Acinetobacter Parvus LUH 4616
T
(NR 025425)
Acinetobacter tandoii CCM 7199
T
(HQ180189)
Acinetobacter Iwoffii DSM 2403
T
(NR 026209)
Acinetobacter brisouii 5YN5-8
T
(DQ832256)
Acinetobacter nosocomialis
RUH 2376
T
(HQ180192)
Acinetobacter calcoaceticus
DSM 30006
T
(AJ633632)
Acinetobacter pittii LMG 1035
T
(HQ180184)
Acinetobacter ursingii LUH 3792
T
(AJ275038)
Acinetobacter bereziniae ATCC 17924
T
(Z93443)
Acinetobacter guillouiae DSM 590
T
(X81659)
Acinetobacter baumannii ATCC 19606
T
(Z93435)
Acinetobacter junii ATCC 17908
T
(Z93435)
Acinetobacter venetianus ATCC 31012
T
(AJ295007)
Acinetobacter towneri CCM 7201
T
(HQ180191)
Acinetobacter baylyi CCM 7195
T
(AM410709)
Acinetobacter rudis G30
T
(EF204258)
SAP 970.1 (JQ771134)
SAP 971.1 (JQ771135)
SAP 220.2 (JQ771129)
SAP 763.2
T
(JQ771132)
SAP 249.1
(JQ771130)
SAP 240.2 (JQ771138)
SAP 239.2 (JQ771137)
SAP 284.1
T
(JQ771141)
Acinetobacter nectaris
SAP 242.2 (JQ771140)
Acinetobacter boissieri
SAP 3 20.1 (JQ771142)
SAP 219.2 (JQ771136)
SAP 241.2 (JQ771139)
Alkanindiges illinoisensis MVAB Hex1
T
(AF513979)
Perlucidibaca piscinae lMCC1704
T
(DQ664237)
Moraxella lawnala ATCC 17967
T
(AF005160)
Ps
y
chrobacter immobilis ATCC 43116
T
(U39399)
SAP 956.2 (JQ771133)
SAP 305.1 (JQ771131)
99
100
98
Acinetobacter radioresistens M 17694
T
(Z93445)
Acinetobacter indicus A648
T
(HM047743)
Acinetobacter soli B1
T
(EU290155)
100
100
100
Acinetobacter gemeri CCM 7197
T
(HQ180188)
100
99
Fig. 1. Neighbour-joining tree, based on 16S rRNA gene sequences, showing the relationships of nectar strains of A. nectaris
sp. nov. and
A. boissieri
sp. nov. with respect to other members of the genus
Acinetobacter
and representatives of closely
related genera within the family Moraxellaceae. Evolutionary distances were computed using the Jukes–Cantor method and are
in the units of the number of base substitutions per site. There were a total of 1320 positions in the final dataset. All positions