




read more










This work advances their understanding of how Nsp15 recognizes and processes viral RNA and will aid in the development of new anti-viral therapeutics. ( which was not certified by peer review ) is the author/funder. This article is a US Government work.
Since the trinucleotide was not bound to the binding site residues during the entire half a microsecond production runs in most systems, the energy calculations were performed for each 50 ns segments (with 50 samples selected at each nanosecond) separately for each trajectory.
other viral proteins may influence Nsp15 RNA targets and regulation in host cells, as Nsp15 is believed to localize within the replication-transcription complex of Nsps, including the RdRp complex (54,55).
Enzymatic activity occurs in the C-terminal EndoU domain, which is more broadly conserved across nidoviruses, suggesting that this endoribonuclease activity is critically important for large, positivestrand RNA viruses (5,6).
Nsp15 is a key player in blocking activation of host dsRNA sensors by preventing the accumulation of viral RNA and a promising therapeutic target (9,13,14).
Work in animals and cell culture has shown that Nsp15 function is not necessary for viral replication, however Nsp15 nuclease activity is critically important for evasion of the host immune response to the virus, specifically by preventing the activation of dsRNA sensors (7- 11).
Rootmean square deviations (RMSDs) were used to establish the stability of the simulated systems (Supplementary Figure 4A) in which the isolated monomer systems displayed elevated dynamics (as assessed by RMSDs) compared to the protomers assembled into the hexamer.
The authors captured the post-cleavage state by incubating Nsp15 with excess unmodified RNA (AUA) prior to vitrification and cryo-EM data collection.
The authors incubated WT Nsp15 with the TRS-N and TRS-S containing RNA substrates and then resolved the cleavage products on denaturing urea gels.
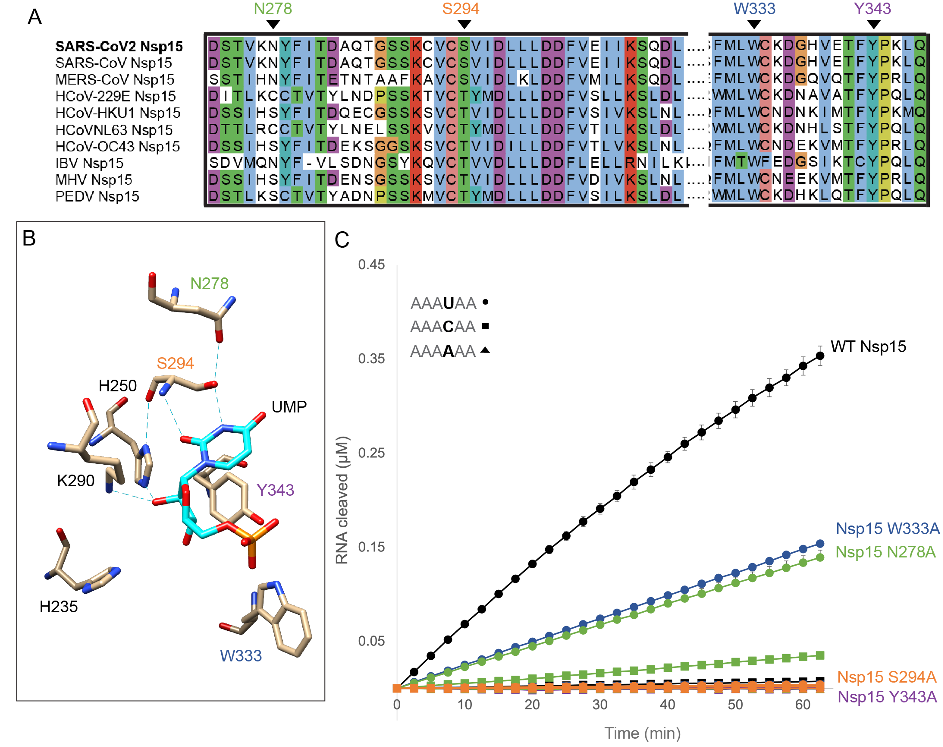
Based on these endoribonuclease assay results, the authors define the consensus motif for Nsp15 cleavage as (N)(U)^(R>U>>C) (where N is any nucleotide and R is a purine).
Consistent with their assay results with a six nucleotide substrate, the N278A Nsp15 variant cleaved more slowly and produced more C cleavage products than WT Nsp15 (Fig. 7).
the authors also looked at Nsp15’s ability to degrade polyU sequences and found that Nsp15 efficiently degrades polyU containing RNAs in vitro.
the authors turned to molecular dynamics simulations to further characterize the behavior of the nucleotides near the active site.