week 2 and, although responses at week 4 were stronger for the ED group, no major differences
were found subsequently. Boost immunization at week 10 increased EC
50
binding titers to the
10
3
-10
4
range for all animals, indicating a strong antibody recall response. ID
50
neutralization titers
showed a similar pattern where both immunization groups developed specific neutralizing
antibody responses by week 4, which were enhanced by week 12, post-boost (figs. S1 and S2).
Next, we examined serum responses for cross-reactivity with SARS-CoV-1 (Fig. 1B, fig. S2).
Strong cross-reactive binding responses to SARS-CoV-1 S protein were observed (Fig. 1B, fig.
S2), as well as strong cross-neutralizing antibody responses against SARS-CoV-1 (Fig. 1C, fig.
S2). ID
50
neutralization titers showed a modest correlation with EC
50
binding titers for both SARS-
CoV-2 and SARS-CoV-1 (Fig. 1D). The elicitation of potent cross-neutralizing responses in rhesus
macaques by S-protein immunization is in stark contrast to human SARS-CoV-2 natural infection,
which typically results in autologous nAb responses where cross-neutralizing activity against
SARS-CoV-1 is rare (Fig. 1E, fig. S3) (34-36). In humans vaccinated with mRNA S-protein, we
similarly found SARS-CoV-2 autologous but not SARS-CoV-1 cross-reactive nAb responses (Fig.
1E, fig. S3) as also described by others (37).
To examine the role of species-specific B cell immunogenetic differences, we vaccinated a group
of 5 mice twice (wk0-prime, wk3-boost) with SARS-CoV-2 S-protein and SMNP adjuvant (fig. S4).
Consistent with earlier studies (38-40), SARS-CoV-2 S protein immunization of mice elicited high
titers of autologous SARS-CoV-2 nAbs; however, the SARS-CoV-1 cross-neutralizing antibody
titers were significantly less than the macaque cross-nAb responses (Fig. 1E, fig. S4), indicating
a strong species-dependent contribution to development of bnAbs to SARS-CoV-2 S protein.
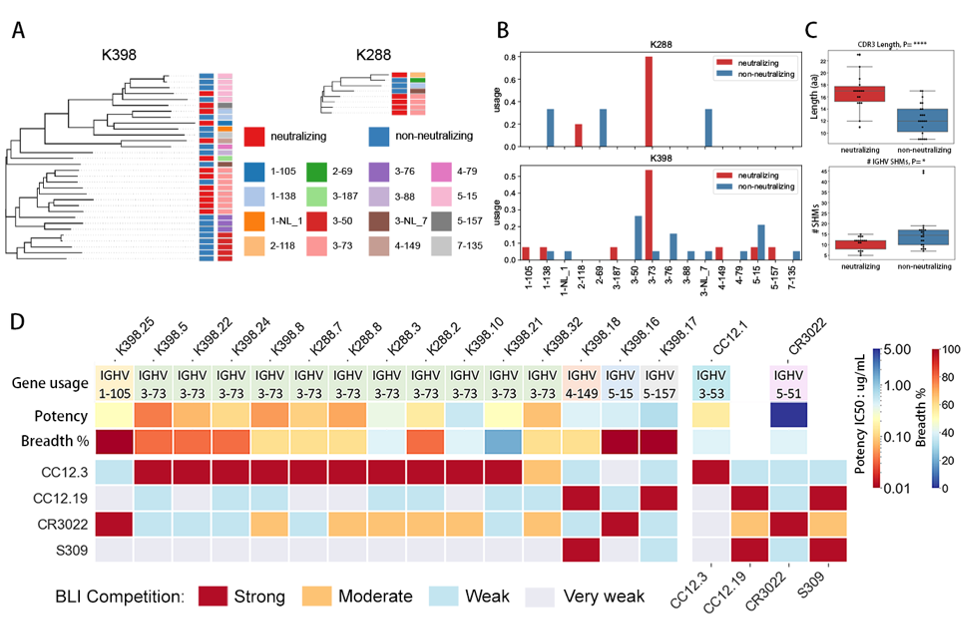
Specificities and breadth of neutralization of the polyclonal macaque nAbs
.CC-BY-NC-ND 4.0 International licenseavailable under a
was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made